Portal Weekly #40: computational pathology, improving AlphaFold2, multi-diffusion, and more
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
💻 Latest Blogs
Last week we had two blog posts! Dominique Beaini introduced MolGPS - a foundation model for molecular property prediction built using GNNs and studying their scaling behaviour.
Frederikke Isa Marin and Felix Teufel introduced BEND, an investigation and benchmarking of the representations learned by DNA language models in an effort to establish biologically-meaningful tasks for the field.
💬 Upcoming talks
M2D2 continues next week with a talk by Chenru Duan, who will explain the development of an object-aware SE(3) equivariant diffusion model that satisfies physical symmetries and constraints in an elementary chemical reaction, and does so in seconds!
Join us live on Zoom on Tuesday, March 26th at 11 am ET. Find more details here.
LoGG continues with a presentation by Peter Eckmann, who will tell us about an approach to achieve optimal trade-off between accuracy and computational cost in generative drug discovery.
Join us live on Zoom on Monday, March 25th at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
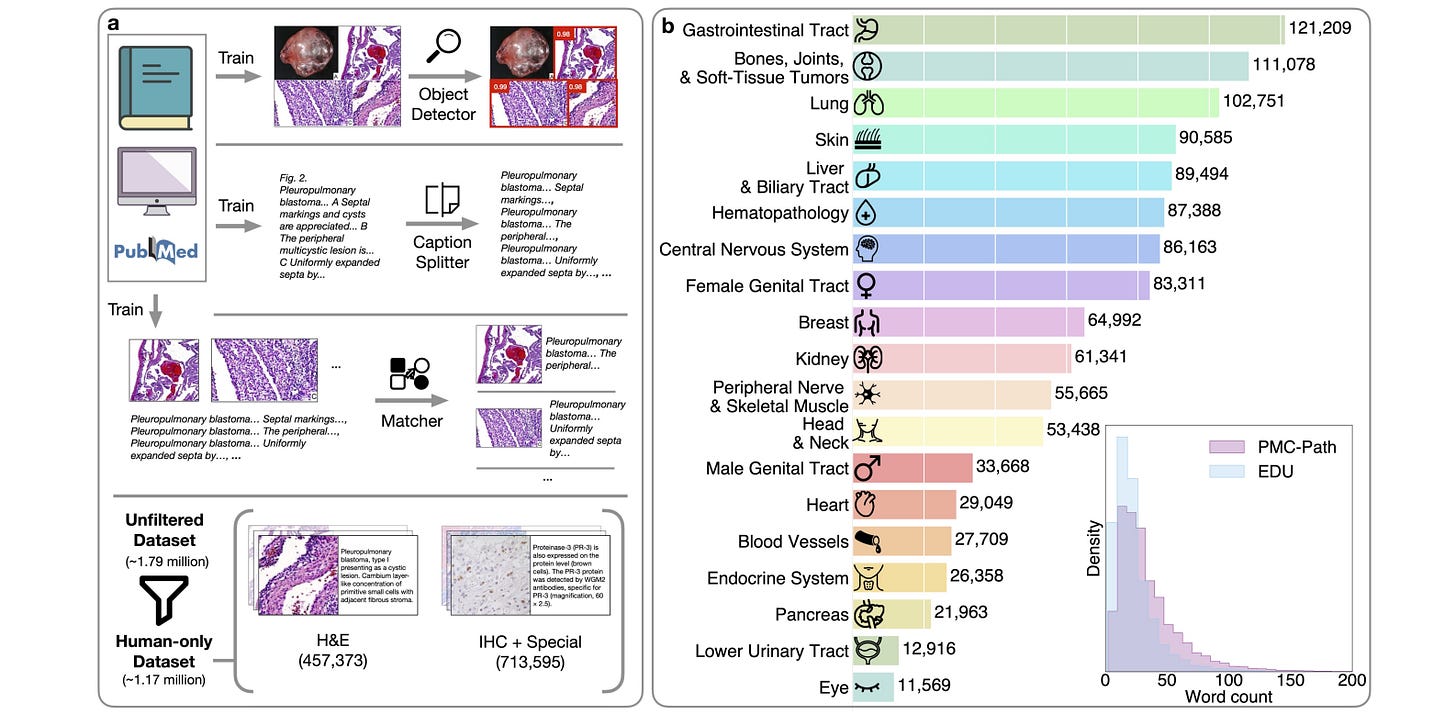
A visual-language foundation model for computational pathology
Automated diagnosis by digital pathology could help accelerate biomedical knowledge and treatment, but training models is often hard because ground-truth labels are sometimes hard to come by and models may not generalize to other tasks or diseases. This study introduces CONCH, a visual-language foundation model that uses histopathology images, biomedical text, and image-caption pairs. Evaluating on 14 diverse benchmarks showed that the model’s knowledge could be transferred to a range of tasks, and its success could inspire other ML-based workflows requiring minimal or no further supervised fine-tuning.
ML for Small Molecules
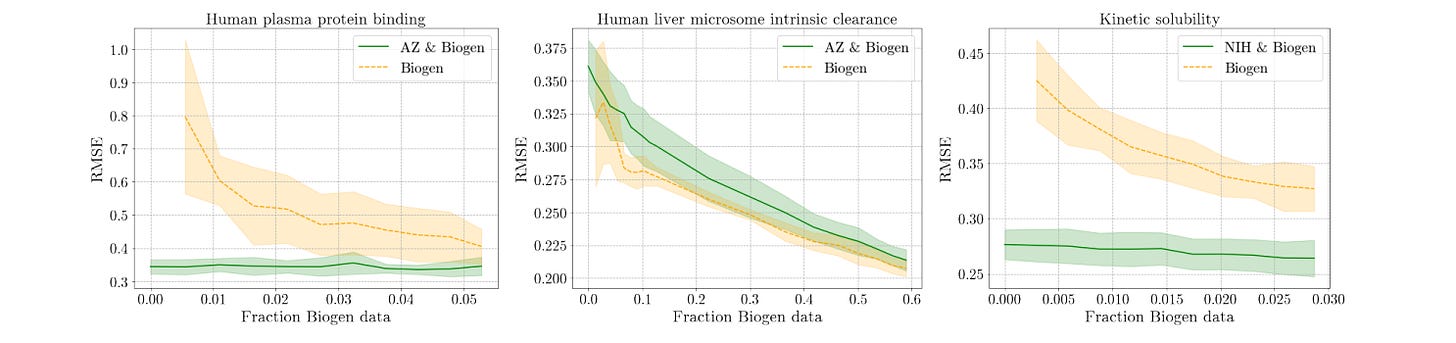
Benchmarking ML in ADMET predictions: the practical impact of feature representations in ligand-based models
When using ML models, the choice of molecule representation can often be critical for performance. Classical techniques often use RDKit descriptors and Morgan fingerprints, or sometimes concatenations between different featurization schemes. This study aims to improve understanding of the impact of feature concatenation, investigate how deep learning representations compare to classical descriptors and fingerprints in ADMET prediction, and integrate cross-validation with statistical hypothesis testing to boost reliability. What are the key takeaways? Read the paper to find out!
ML for Proteins
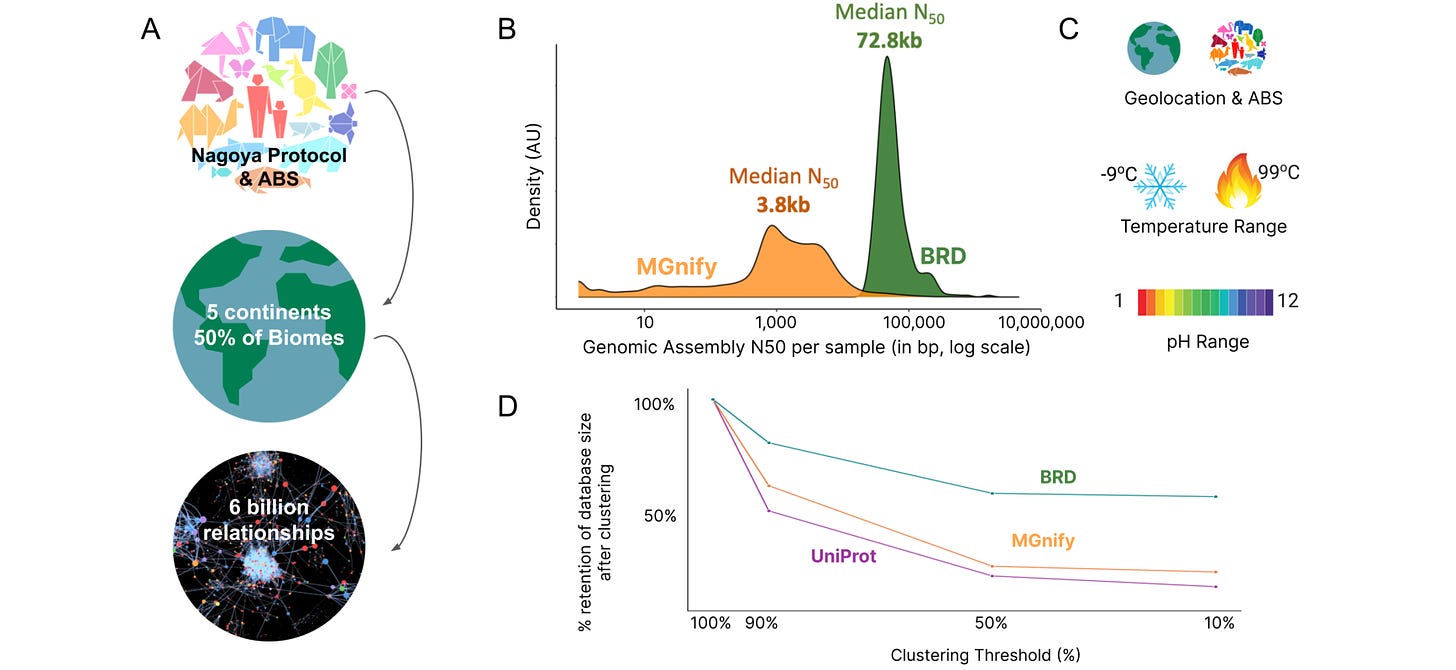
Improving AlphaFold2 Performance With a Global Metagenomic & Biological Data Supply Chain
You may think that we have a lot of DNA and protein sequence information, and in some ways, it’s true: the number of sequences in our databases has exploded. However, it’s suspected that we’ve only sequenced a tiny fraction of worldwide biodiversity, which could mean that we’re missing some useful proteins and small molecules that could be used as treatments and for other purposes. Basecamp research partnered with nature parks and biodiversity stakeholders across 5 continents (and covering 50% of global biomes) to establish a diverse global metagenomics dataset. Supplementing the obtained multiple sequence alignments (MSAs) during inference of AlphaFold2, they seem to get improved structure predictions and docking results, showing that diversity is valuable.
ML for Omics
Geometric deep learning framework for de novo genome assembly
We’re getting a huge number of genome sequences these days, but reconstructing genomes from a bag of individual reads is not trivial, especially with repetitive regions of DNA. This study presents GNNome, a graph neural network framework that leverages symmetries inherent to the problem. Their results suggest that the approach has similar or better performance with state-of-the-art tools across several different species. With more assembled genomes at our disposal in the future, the model has even more room to improve. The authors also provide the framework and best-performing model as a tool that you can use to assemble new haploid genomes and as a basis for improvement in this domain.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
Evaluating retrosynthesis with synthesis and retro-fallback by Austin Tripp
LoGG
MultiDiffusion: Fusing Diffusion Paths for Controlled Image Generation by Omer Bar-Tal
CARE
Additive Decoders for Latent Variables Identification and Cartesian-Product Extrapolation by Sébastien Lachapelle
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋