Portal Weekly #43: combating antibiotic resistance, investigating invalid SMILES, and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
💬 Upcoming talks
M2D2 continues next week with a talk by Minsu Kim, who will present a new training algorithm designed to enhance the sampling quality of GFlowNets.
Join us live on Zoom on Tuesday, April 16th at 11 am ET. Find more details here.
LoGG continues with a presentation by Lior Yariv, who will introduce a simple 3D shape representation that allows efficient conversion of large 3D datasets to representation form, provides a good tradeoff of approximation power vs. number of parameters, and has a simple tensorial form compatible with existing neural architectures.
Join us live on Zoom on Monday, April 15th at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter!
💻 Upcoming Events
We are excited to present the inaugural machine learning for drug discovery summer school happening at Mila from June 12th to 21st! If you have experience working with AI (i.e. data scientist, ML engineer, etc.) and you’re interested in applying your skills to the world of drug discovery, this is perfect for you. This will also be a great opportunity for scientists (biochemists, biologists, chemists etc.) with programming skills looking to expand their experience in ML for drug discovery.
Registration is now open! Find more details here.
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
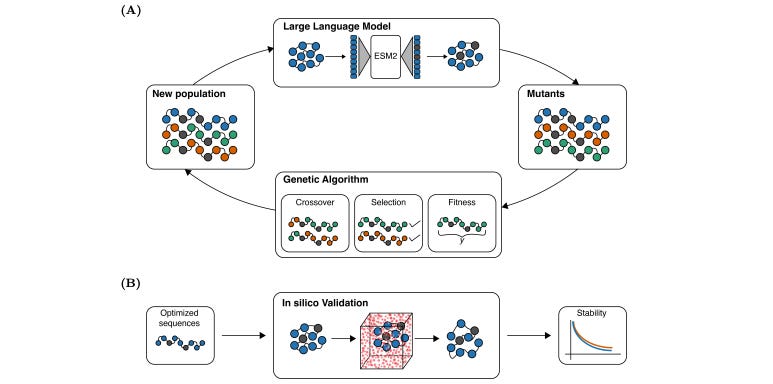
Integrating Genetic Algorithms and Language Models for Enhanced Enzyme Design
LLMs are powerful tools for modeling and analyzing biological sequences, but their application to protein design is limited by the high combinatorial complexity of proteins. This study introduces a framework that combines LLMs with genetic algorithms (GAs) to optimize enzymes. As Austin Tripp explained in his blog post, genetic algorithms are an older (but very useful) technique for iterating on versions of the best molecules encountered so far. These researchers find that combining LLMS and GAs to improve enzyme reactions and turnover rates potentially gives “the best of both worlds”, with generated mutants outperforming wild-type enzymes in 90% of cases. Now to test those samples in the lab.
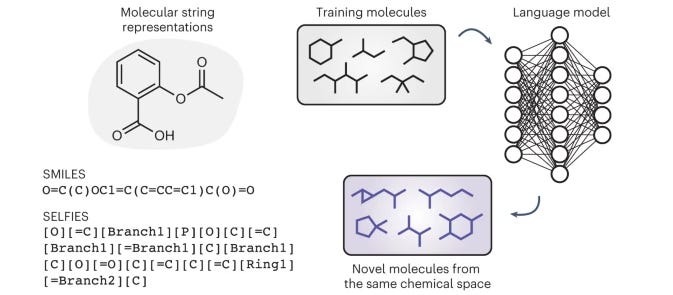
Invalid SMILES are Beneficial Rather Than Detrimental to Chemical Language Models
Language models trained on SMILES (Simplified Molecular-Input Line-Entry System) representations show great promise for molecule generation (check out talks by Amir Barati Farimani and Rıza Özçelik!) but generate some number of invalid SMILES that can’t be decoded to a chemical structure. This seems like a flaw, and work is going into ameliorating the problem. This paper argues that the ability to produce invalid outputs is not harmful but is instead beneficial to chemical language models, and that enforcing valid outputs produces structural biases in the generated molecules, impairing distribution learning and limiting generalization to unseen chemical space. This casts chemical language models in a different light, and could spur additional research on this topic.
ML for Small Molecules
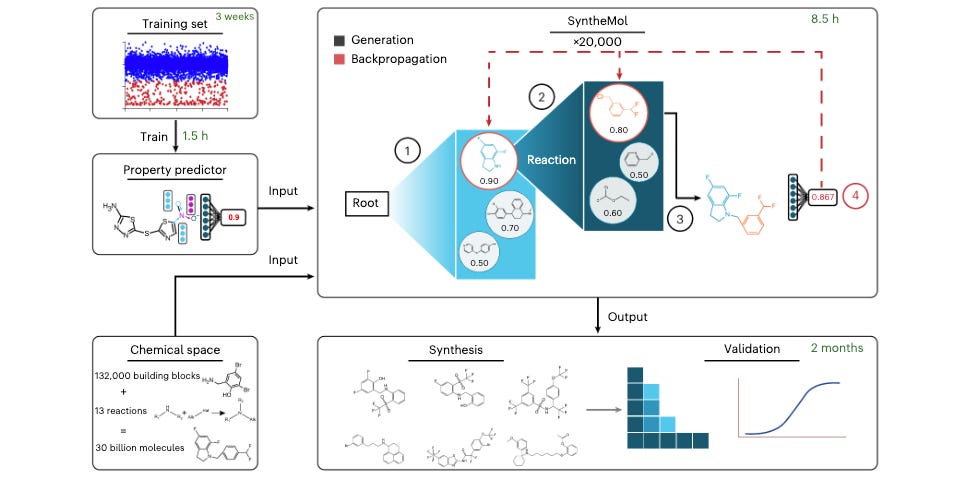
Generative AI for Designing and Validating Easily Synthesizable and Structurally Novel Antibiotics
Antibiotic resistance in bacteria is a critical health concern. As our current antibiotics stop working, if we have nothing to replace them with it will not only make infections harder to treat but other medical procedures like surgery and cancer chemotherapy much riskier. This study presents SyntheMol, a generative model that designs new antibiotic compounds that are easy to synthesize out of the total vast chemical space. Out of the 58 synthesized molecules they generated, six showed antibacterial activity against ESKAPE pathogens, which are especially virulent and drug-resistant. This well-considered study shows the potential of ML to address critical problems, with their results nicely supported by experimental validation.
AI is a Viable Alternative to High Throughput Screening: a 318-Target Study
High-throughput screening (HTS) is routinely used but has physical limitations, which limits the chemical space that can be searched. This study describes the largest and most diverse virtual HTS campaign reported to date, with 318 individual projects together finding novel hits across every major therapeutic area and protein class. Molecules found this way were novel scaffolds rather than minor modifications to known bioactive compounds. This study represents practical progress for graph convolution networks (which their AtomNet model is based on) and for in silico screening.
ML for Atomistic Simulations
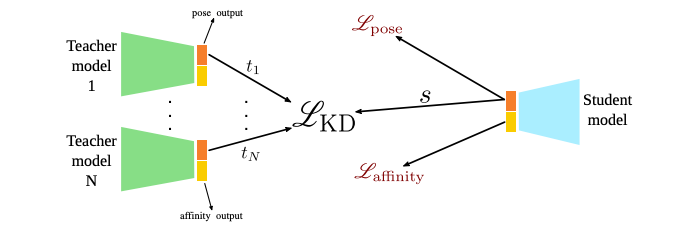
Condensing Molecular Docking CNNs via Knowledge Distillation
Some work has suggested that convolutional neural networks have potential in molecular docking scoring, but they suffer from heavy computation. An ensemble of CNN models can get better performance than a single model but requires even more compute. This study aims to use knowledge distillation (KD) to reduce the computational overhead of using multiple CNNs. Their investigation suggests that this approach produces more powerful single models than those without KD, with the distilled model’s performance related to the pose score variation within the teacher ensemble.
ML for Proteins
Learnt Representations of Proteins Can be Used for Accurate Prediction of Small Molecule Binding Sites on Experimentally Determined and Predicted Protein Structures
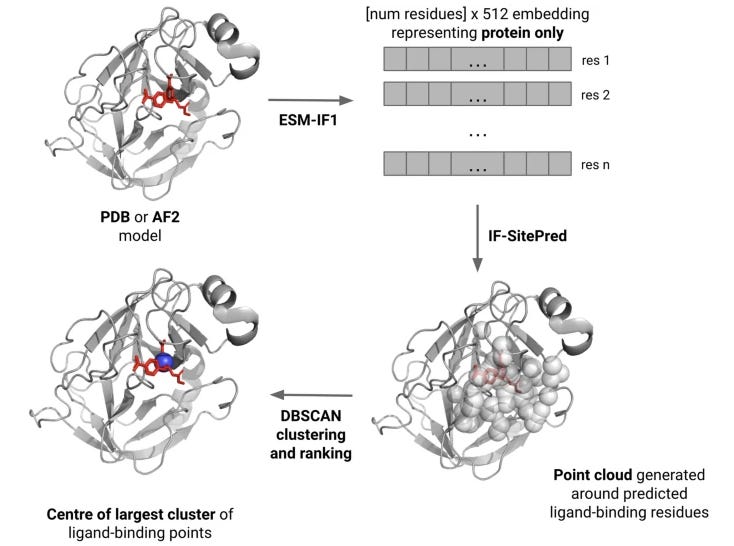
In early drug development we usually want to identify potential ligand-binding sites in a protein; there are many methods to do so but they have been designed for use on experimentally-determined structures. When we don’t have an experimental structure for the protein we’re interested in, we could use computationally predicted structures instead but this is often considered “risky” because getting things wrong can be costly. This study describes a protein-ligand binding site prediction method based on labeling protein language model embeddings. Their results suggest that their IF-SitePred model is competitive with methods that predict binding sites on experimental structures, but that it can also do better on novel proteins where low accuracy has been simulated by molecular dynamics. If more ‘classical’ techniques would fail, this may be a reasonable next bet.
ML for Omics
Genomic Language Model Predicts Protein Co-Regulation and Function
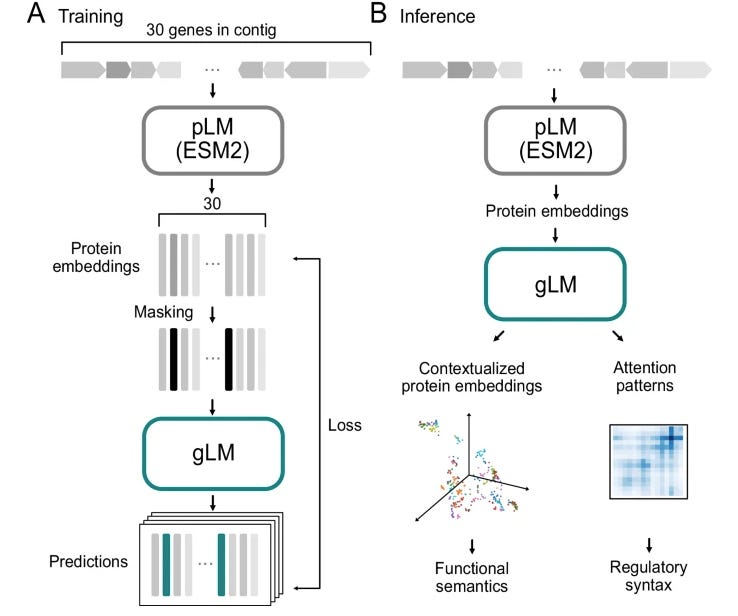
The genome is a complex place, and context can matter a lot. This study describes a genomic language model trained on millions of metagenomic scaffolds (bacterial genome sequences reconstructed from environment samples) to try to capture higher order genomic context information. Their analyses suggests that the model learns co-regulated functional modules (operons, groups of genes that are activated together in bacteria), without using co-expression cues that would be given from transcriptomics. This suggests that the model is learning a certain amount of complex relationships and functional semantics of genes in their genomic regions.
The Impact of Package Selection and Versioning on Single-Cell RNA-Seq Analysis
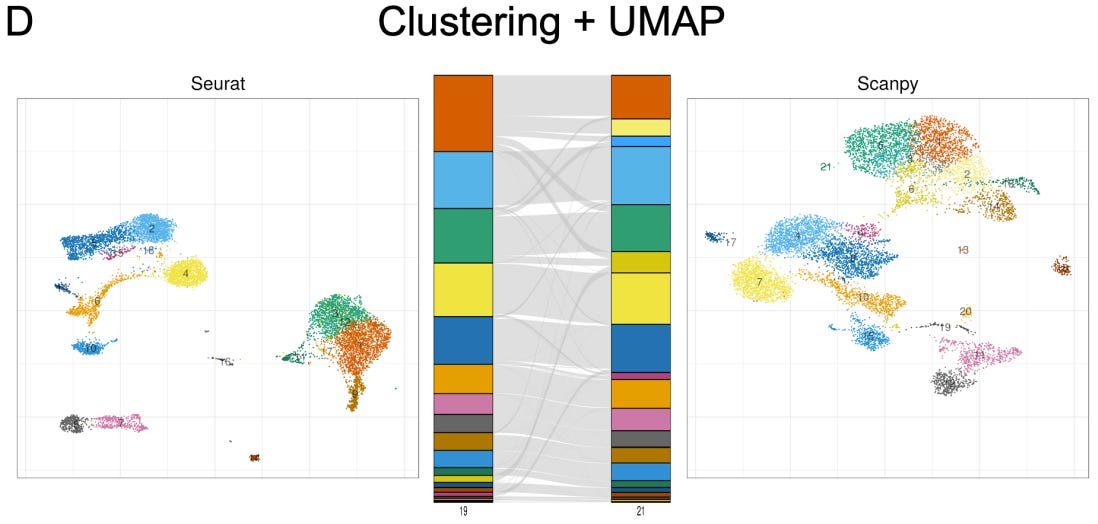
In this article and a Twitter thread (recommended for a digestible breakdown of the results), the authors compare the two dominant packages for implementing single-cell RNA-seq workflows, finding that there are considerable difference between the outputs of the two. The extent of differences between the programs is roughly equivalent to the variability that would be introduced by sequencing less than 5% of the reads for scRNA-seq experiments, or by analyzing less than 20% of the cell population. This is huge, and highlights that those developing scientific software need to prioritize transparency, consistency, and reproducibility for their tools, while users should carefully assess tools upon which they’re relying.
Open Source
DrugDomain: the Evolutionary Context of Drugs and Small Molecules Bound to Domains
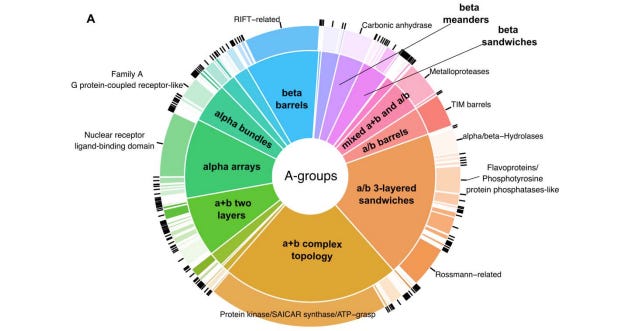
The Evolutionary Classification Of protein Domains (ECOD) classifies domains into an evolutionary hierarchy that focuses on distant homology. Previously, no structure-based protein domain classification existed that included information about both the interaction between small molecules or drugs and the structural domains of a target protein. This preprint describes a database that reports the interaction between ECOD domains of human target proteins and DrugBank molecules and drugs.
Reviews
Membrane Transporters in Drug Development and as Determinants of Precision Medicine
Membrane transporters can affect drug efficacy in a number of ways, from actively letting drugs into cells (which we might want) to pumping them out (which we probably don’t want). If you want to understand more about the biology behind this important group of proteins and how it can affect drug development efforts, check out this review.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
Combinatorial prediction of therapeutic perturbations using causally-inspired neural networks by Guadalupe Gonzalez
LoGG
Stability-Aware Training of Neural Network Interatomic Potentials with Differentiable Boltzmann Estimators by Sanjeev Raja
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋