Portal Weekly #44: evaluating if LLMs are superhuman chemists, Foldseek-Multimer, and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
💻 Latest Blogs
This week we had a blog post by Tony Shen, who argues that just using a maximum likelihood estimation objective is not sufficient for diffusion models to make good structures in drug design. See his proposed solution here.
Interested in sharing your work with the community? Fill out our interest form to get started.
💬 Upcoming talks
M2D2 continues next week with a talk by Tinglin Huang, who will present MolGroup, a strategy for identifying which auxiliary datasets we can include that won’t contradict the knowledge in our target dataset.
Join us live on Zoom on Tuesday, April 23rd at 11 am ET. Find more details here.
LoGG continues with a presentation by Sergey Pozdnyakov, who will present a general symmetrization method that adds rotational equivariance to any given point-cloud model while preserving all the other requirements.
Join us live on Zoom on Monday, April 22nd at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter and LinkedIn!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
Are Large Language Models Superhuman Chemists?
LLMs have emergent properties - they can perform tasks on which they have not been explicitly trained but we don’t really know exactly how they do so. We only have a limited understanding of the chemical reasoning abilities of LLMs, which we need to be able to design more effective models. This paper introduces "ChemBench," an automated framework designed to rigorously evaluate the chemical knowledge and reasoning abilities of state-of-the-art LLMs against the expertise of human chemists. They have curated more than 7,000 question-answer pairs for a wide array of subfields of the chemical sciences, evaluated leading open and closed-source LLMs, and found that the best models outperformed the best human chemists in our study on average. Their findings highlight the importance of continuing to develop evaluation frameworks to produce safe and useful LLMs.
ML for Small Molecules
MolSnapper: Conditioning Diffusion for Structure Based Drug Design
There are different ways to generate a molecule against a specific target. If you have a protein structure, you can try to generate a 3D molecule within the target binding site Autoregressive models do this by iteratively adding atoms and bonds but can accumulate errors and can lack global context. Diffusion-based models can model local and global interactions but have been trained on a general molecular space, not a target pocket. This study proposes using a pretrained diffusion model but the generative process on 3D structural information with pharmacophores. Their results suggest that their approach gives more valid molecules that bind well to a given binding site than other methods.
Is Meta-training Really Necessary for Molecular Few-Shot Learning ?
It’s expensive to produce data points in drug discovery, so a field of study has sprung up to try to provide reliable predictions while using a small amount of labelled data. Meta-learning, using methods to “learn to learn” from a small set of labelled examples, is a solution that some argue is the default and best solution. This study suggests that a simple fine-tuning approach is highly competitive with meta-learning in this space. They introduce a new benchmark to assess the robustness of the competing methods to domain shifts and find that their approach consistently gets better results than meta-learning methods.
ML for Atomistic Simulations
FABind+: Enhancing Molecular Docking Through Improved Pocket Prediction and Pose Generation
At NeurIPS 2023, a model called FABind for end-to-end pocket prediction and docking was proposed; combining pocket prediction and docking was suggested to be able to speed up the model. Building on the previous work, the authors now propose FABind+, a version that they suggest largely boosts the performance of its predecessor. Their results suggest competitive performance with other state-of-the-art models while being remarkably fast.
This paper uses a different benchmark than PoseBusters - watch Martin Buttenschoen’s talk to see why that may be an issue.
ML for Proteins
Rapid and Sensitive Protein Complex Alignment with Foldseek-Multimer
We have access to millions of predicted protein structures with AlphaFold DB and the ESM Metagenomic Atlas, but searching these databases can be a bottleneck. Foldseek was developed to address this, aligning the structure of a query protein against the database while decreasing computation times compared to other methods. This paper presents an extension to Foldseek, allowing search on a protein complex level. This is very useful since many proteins don’t operate alone but as complexes. The authors suggest that Foldseek-Multimer is 3-4x faster than the gold standard while producing comparable results, allowing for the comparison of billions of complex pairs in a day.
ML for Omics
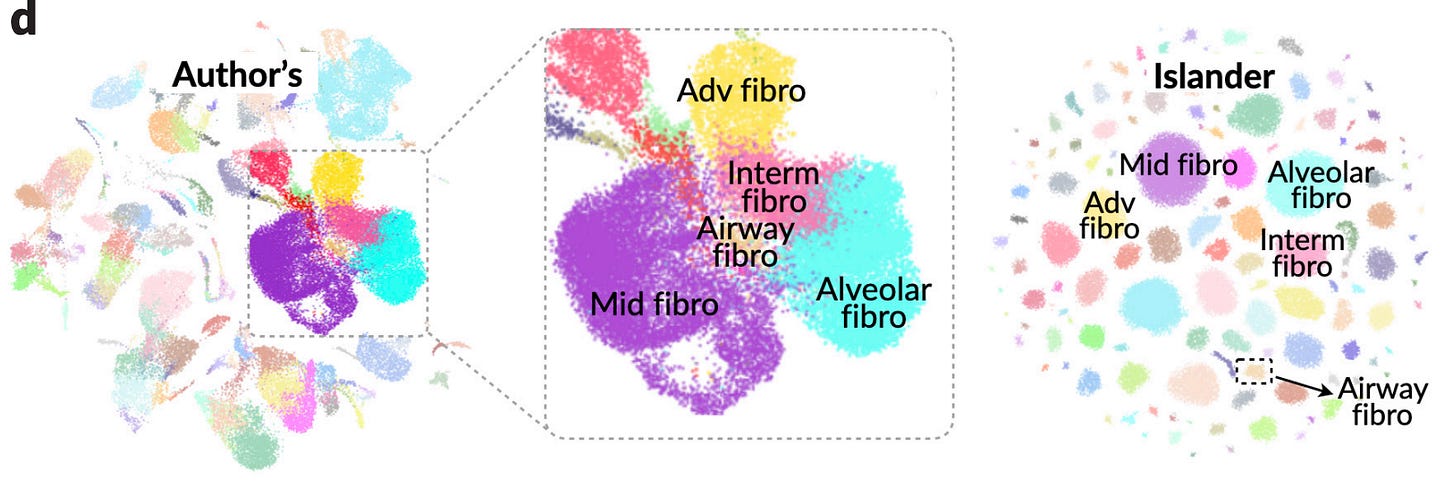
Metric Mirages in Cell Embeddings
Biological studies increasingly rely on embeddings of single-cell profiles, but it can be hard to assess the quality of these embeddings. We need good ways of making such assessments to avoid misleading biological interpretations, assessing integration methods, and establishing the zero-shot capabilities of foundational models. This study suggests that current evaluation metrics can be highly misleading, and train a three-layer perceptron, Islander , which outperforms all 11 leading embedding methods on a diverse set of cell atlases, but in fact distorts biological structures. They also suggest a metric, scGraph, to flag these kind of distortions.
Reviews
Deep Learning for Molecules and Materials
Deep learning is becoming a standard tool in chemistry and materials science. This book presents deep learning as a set of tools that provide powerful feature engineering and modeling capability. It is aimed towards students with a programming and chemistry background that are interested in building competency in deep learning.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
Local Search GFlowNets by Minsu Kim
LoGG
Mosaic-SDF for 3D Generative Models by Lior Yariv
CARE
Uncovering and Inducing Interpretable Causal Structure in Deep Learning Models by Atticus Geiger
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋