Portal Weekly #45: ICLR TechBio social, optimal transport for chemical transition states, alchemical MLIPs, designing gene editors, and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
This week, we hit 1,000+ members on Portal! 🎉
Thank you to everyone for playing a role in helping us grow and foster the TechBio community. We hope to continue bridging the gap between AI/ML and drug discovery through our reading groups, blogs, events, and more. If you’re not yet on Portal, consider joining - you can participate in discussion forums, DM other community members, and more.
📅 Upcoming Events
Who’s going to ICLR in a few weeks?? Valence Labs is hosting a TechBio social on the evening of May 8th. It’ll be a great opportunity to meet more folks in the community, we hope to see many of you there. Spots are limited so RSVP here.
💬 Upcoming talks
M2D2 continues next week with a talk by Puck van Gerwen, who will introduce EquiReact, an equivariant neural network to infer properties of chemical reactions, built from three-dimensional structures of reactants and products.
Join us live on Zoom on Tuesday, April 30th at 11 am ET. Find more details here.
LoGG continues with a presentation by Farhan Khodaee, who will present in integrated genetics framework to analyze and interpret the high-dimensional landscape of genotypes and their associated phenotypes simultaneously, and a multimodal foundation model built on this framework.
Join us live on Zoom on Monday, April 29th at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter and LinkedIn!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
Bioinformatics Copilot 1.0: A Large Language Model-Powered Software for the Analysis of Transcriptomic Data
Single-cell transcriptomics has exploded in the past decade (a nice technology feature was recently published in Nature Methods), but the volume of data generated means it takes a long time to analyze and many steps. This work presents an LLM-powered software to help users analyze data through a natural language interface without needing proficiency in programming languages. This may help to speed up data analysis and accelerate advancements in the biomedical sciences.
Inferring the Phylogeny of Large Language Models and Predicting Their Performances in Benchmarks
If you know how similar one LLM is to another, can you predict its performance? This study applies phylogenetic algorithms to a set of 77 open-source and 22 closed models, constructing a family tree that captures different LLM families. They suggest that this approach enables a time and cost-effective estimation of LLM capabilities, even without transparent training information.
ML for Small Molecules
React-OT: Optimal Transport for Generating Transition State in Chemical Reactions
Transition states (TS) of molecules can help us understand reaction mechanisms and design better catalysts but are hard to capture in experiments. This study presents an optimal transport approach for generating unique TS structures from reactants and products. The model makes accurate TS structures with RMSD 0.053Å, with fast inference.
First author Chenru Duan gave an M2D2 talk on the subject before this preprint came out - check it out!
Transformers for Molecular Property Prediction: Lessons Learned From the Past Five Years
Transformer models are powerful in many domains, but how are they used in molecular property prediction (MPP)? This review analyzes the currently available models and explores key questions that arise when training and fine-tuning a transformer model for MPP; they also address challenges in comparing different models, and highlight areas not yet covered in current research.
If you’re interested in how transformers (and LLMs) are being used in other aspects of chemistry and drug discovery, check out Andres M Bran’s blog post!
ML for Atomistic Simulations
Interpolation and Differentiation of Alchemical Degrees of Freedom in Machine Learning Interatomic Potentials
Machine learning interatomic potentials (MLIPs) have become a workhorse of modern atomistic simulations, but their computational cost limits their applicability to all problems. This study exploits the fact that graph neural network MLIPs represent discrete elements as real-valued tensors, allowing continuous and differentiable alchemical degrees of freedom in atomistic materials simulations.
If you want to know more about training neural network potentials for MD simulations, start with Stephan Thaler’s M2D2 talk!
ML for Proteins
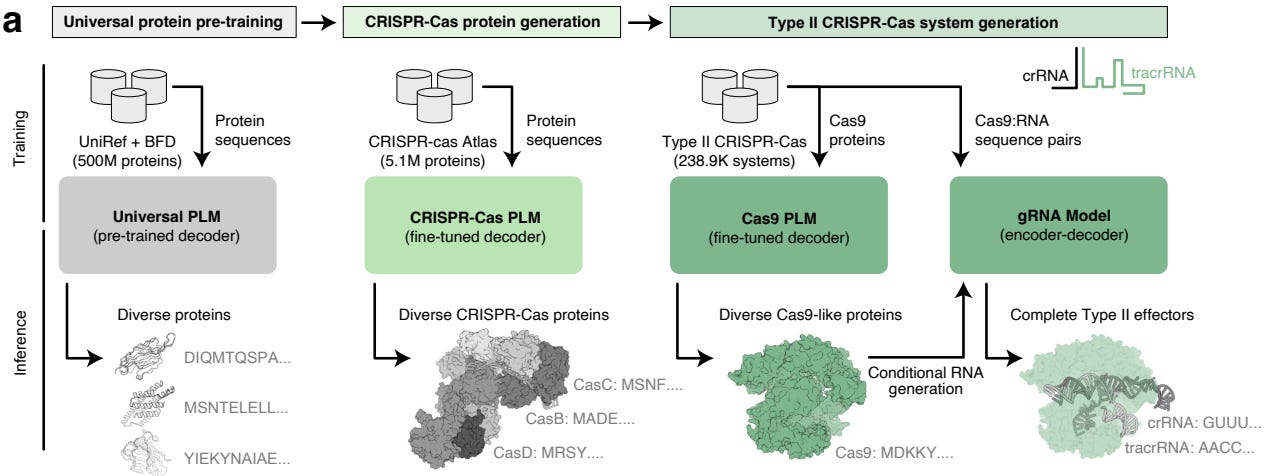
Design of Highly Functional Genome Editors by Modeling the Universe of CRISPR-Cas Sequences
CRISPR-based gene editing has revolutionized biotechnology and research, but proteins that evolved for use in microbes can have undesirable off-target effects when used in different systems. This study curates a dataset of over 1 million CRISPR operons and uses it to train an LLM to generate new proteins that have improved activity or specificity. The work includes experimental characterization, and they also released a publicly available AI-generated gene editor called OpenCRISPR-1.
ML for Omics
Nicheformer: a Foundation Model for Single-Cell and Spatial Omics
Foundation models are trained on large amounts of data with the idea that robust representations can be applied to other tasks. This study aims to exploit a niche in the single-cell foundation model landscape, proposing to generalize recent foundation modeling approaches for disassociated single-cell transcriptomics to the spatial omics setting, using more than 110 million cells in a partial spatial context.
Open Source
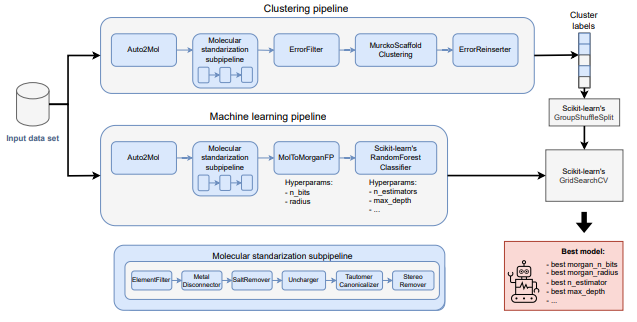
MolPipeline : A Python Package for Processing Molecules With RDKit in Scikit-Learn
Scikit-learn’s Pipeline class allows users to join data transformation steps to an ML model. This paper extends the concept to bioinformatics by wrapping RDKit functionalities like reading and writing SMILES strings and calculating molecular descriptors, as well as common tasks like scaffold splitting and molecule standardization. This could be useful for quickly prototyping strategies end-to-end.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
Learning to Group Auxiliary Datasets for Molecule by Tinglin Huang
LoGG
Smooth, exact rotational symmetrization for deep learning on point clouds by Sergey Pozdnyakov
CARE
Causal Abstractions using Generalized Functions by Sander Beckers
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋