Portal Weekly #48: MISATO, diffusing and denoising drug data, indexing life's biological sequences, and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
📅 Upcoming Events
We have a few tickets left for this years Molecular Machine Learning Conference taking place on June 19th at Mila. After all the news surrounding the AlphaFold 3 release, we’re excited to hear what Max might have to say at his keynote talk at MoML. 👀
Register for a ticket today to hear from Max, Jian Tang, and other speakers!
💻 Latest Blogs
In a blog post this week, Nima Shoghi JMPs into foundation models for chemistry with Joint Multi-Domain Pre-Training, a GNN model trained on diverse datasets spanning different chemical domains that beats or matches the state-of-the-art on 34/40 benchmarks. Check it out here.
💬 Upcoming talks
LoGG continues next week with a talk by Snir Hordan, who will introduce WeLNet architecture, which can process position-velocity pairs, compute functions fully equivariant to permutations and rigid motions, and is provably complete and universal.
Join us live on Zoom on Monday, May 6th at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter and LinkedIn!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
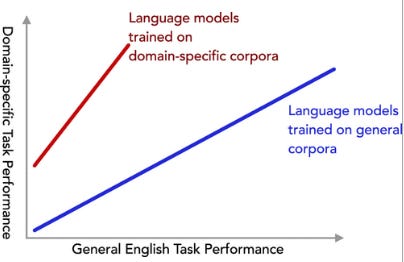
How Beneficial is Pretraining on a Narrow Domain-Specific Corpus for Information Extraction About Photocatalytic Water Splitting?
This study examines how the specificity of training data impacts the performance of language models on specialized tasks. Researchers tested models trained on corpora (of varying levels of specificity) for extracting information on photocatalytic water splitting and found that more specific training data improved performance. Notably, a model trained exclusively on scientific papers about photocatalytic water splitting outperformed previous models, showing significant gains in precision and recall.
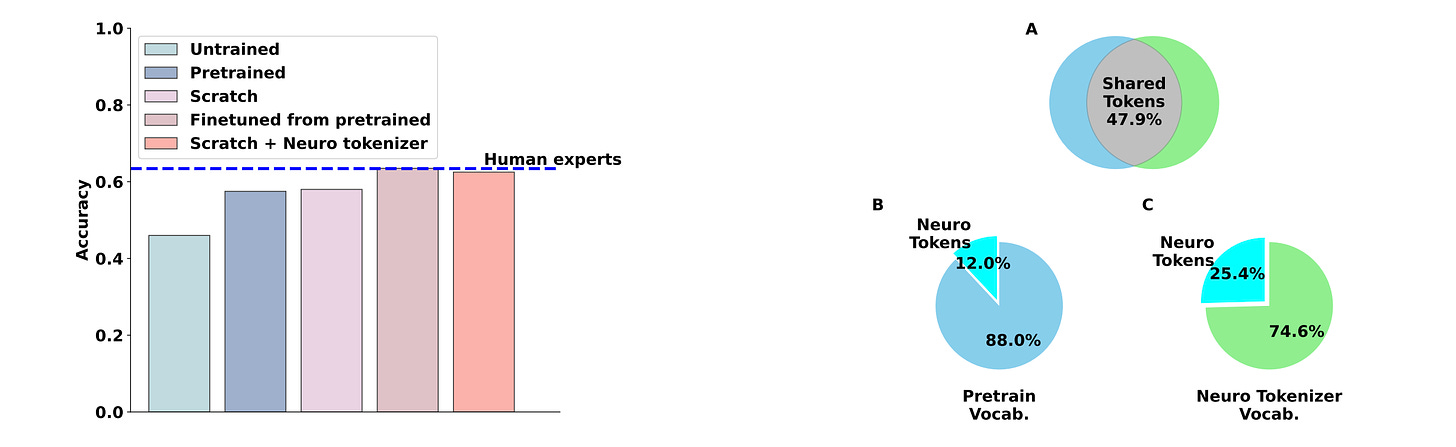
Matching Domain Experts by Training From Scratch on Domain Knowledge
LLMs have outperformed human experts in predicting the results of neuroscience experiments. This study investigates the possibility that statistical patterns in that specific scientific literature underlie LLMs’ performance, rather than emergent reasoning abilities arising from broader training. Their results suggest that at least for neuroscience, small models trained on domain-specific information are sufficient to match expert performance.
ML for Small Molecules
Denoising Drug Discovery Data for Improved ADMET Property Prediction
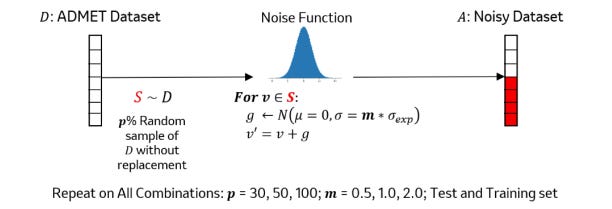
In drug discovery, experimental error inherent in the data affects models’ ability to predict ADMET properties. This study applies deep learning-based data denoising methods to improve ADMET models, focusing on regression tasks rather than classification. Their work suggests that using training error to identify noise in the data leads to significant performance improvements, potentially benefiting models for other experimental assay data.
Synthetic Data from Diffusion Models Improve Drug Discovery Prediction
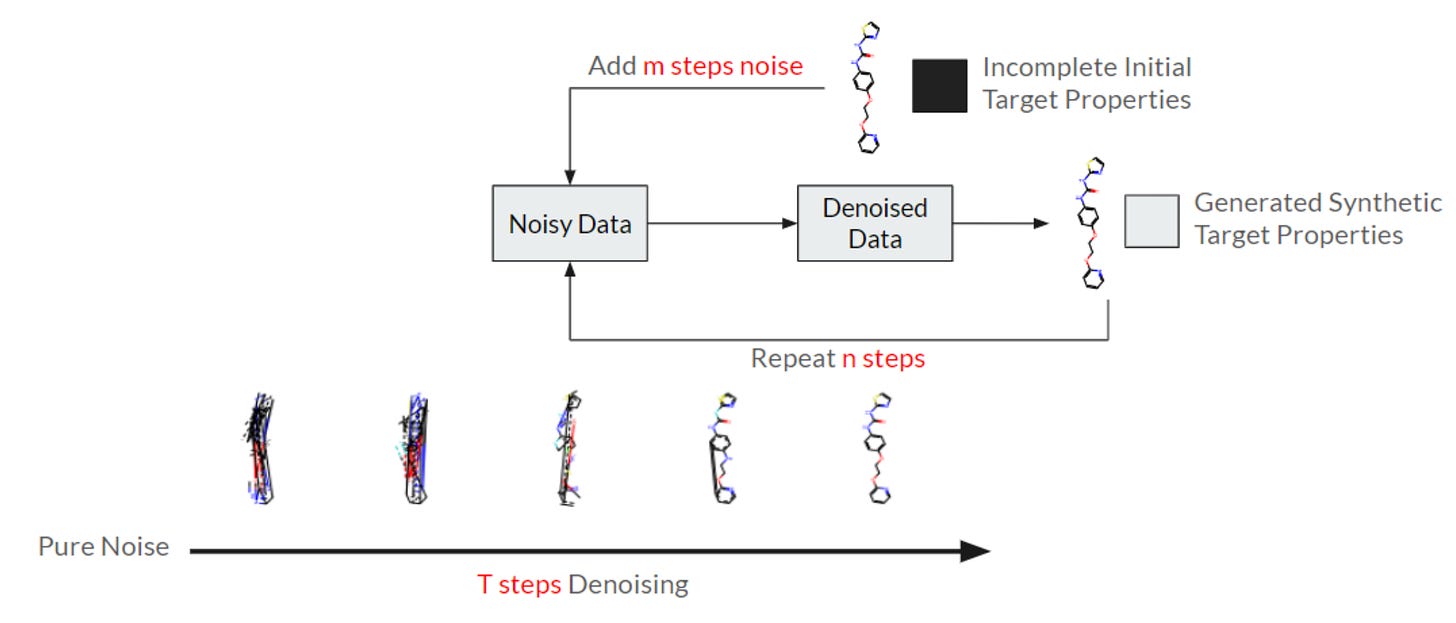
Datasets are often collected independently from each other, often with little overlap, creating data sparsity and making curation difficult. This study proposes a new diffusion GNN model called Syngand that can generate ligand and pharmacokinetic data end-to-end, which can help to ‘fill in the blanks’. Their initial results suggest that this synthetic data helps models to predict downstream molecular properties, although more work is needed.
Optimal Molecular Design: Generative Active Learning Combining REINVENT with Absolute Binding Free Energy Simulations
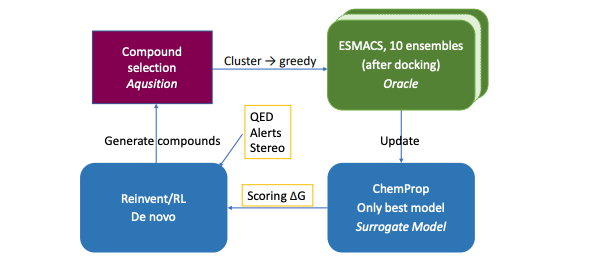
Active learning (AL) uses machine learning to smartly select the next molecular structures to evaluate, similar to the iterative process of lab experiments for optimizing compounds. This study describes a new AL protocol combining AI-driven molecule generation (REINVENT) and physics-based simulations (ESMACS) to discover new ligands for two target proteins, using the powerful Frontier supercomputer. Their findings suggest this protocol can identify better and more chemically diverse binders than traditional methods.
ML for Atomistic Simulations
MISATO: Machine Learning Dataset of Protein–Ligand Complexes for Structure-Based Drug Discovery
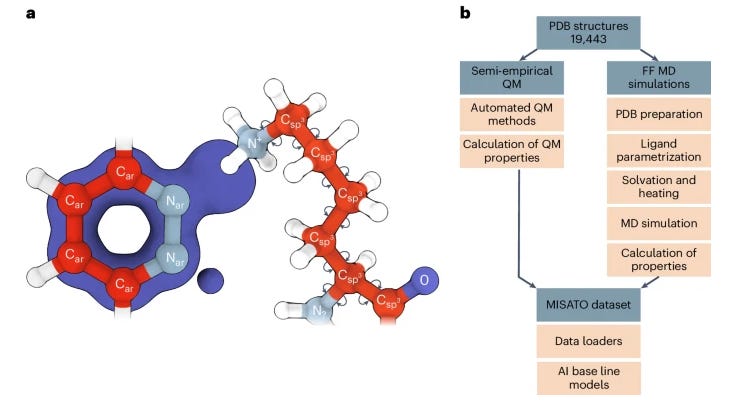
Precise biomolecule–ligand interaction datasets are urgently needed for large language models to help make gains in structure-based drug discovery, quantum chemistry and structural biology. This paper presents MISATO, a dataset that combines quantum mechanical properties of small molecules with molecular dynamics simulations of around 20,000 experimental protein–ligand complexes, validated extensively against experimental data. The dataset includes refined structures and a substantial collection of molecular dynamics traces, which early examples suggest can enhance machine learning models' accuracy, providing a potentially valuable resource for advancing AI-driven drug discovery.
ML for Proteins
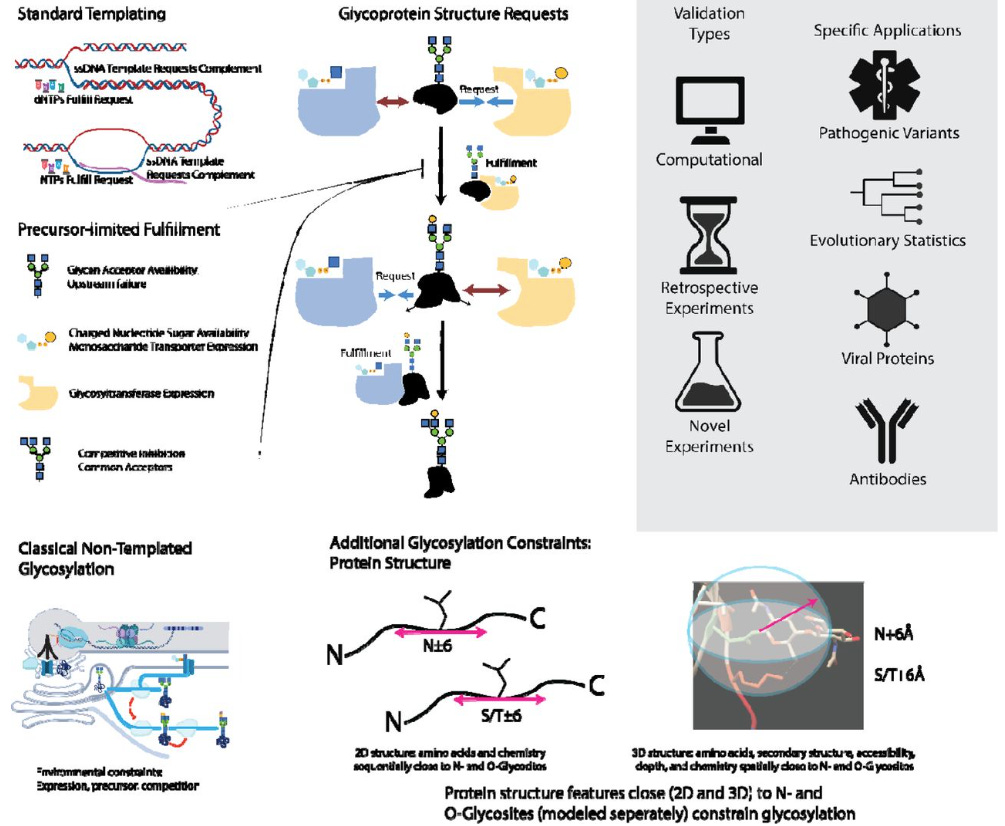
Protein Structure, a Genetic Encoding for Glycosylation
Glycosylation, unlike DNA, RNA, and protein synthesis, is thought to be unconstrained by a template. This study identifies protein-sequence and structural features predictive of glycan patterns and develops a "glycoimpact" matrix to quantify these relationships. Their findings, validated across various proteins, suggest glycan biosynthesis follows specific genetic rules, contradicting the idea of template-free glycosylation.
ML for Omics
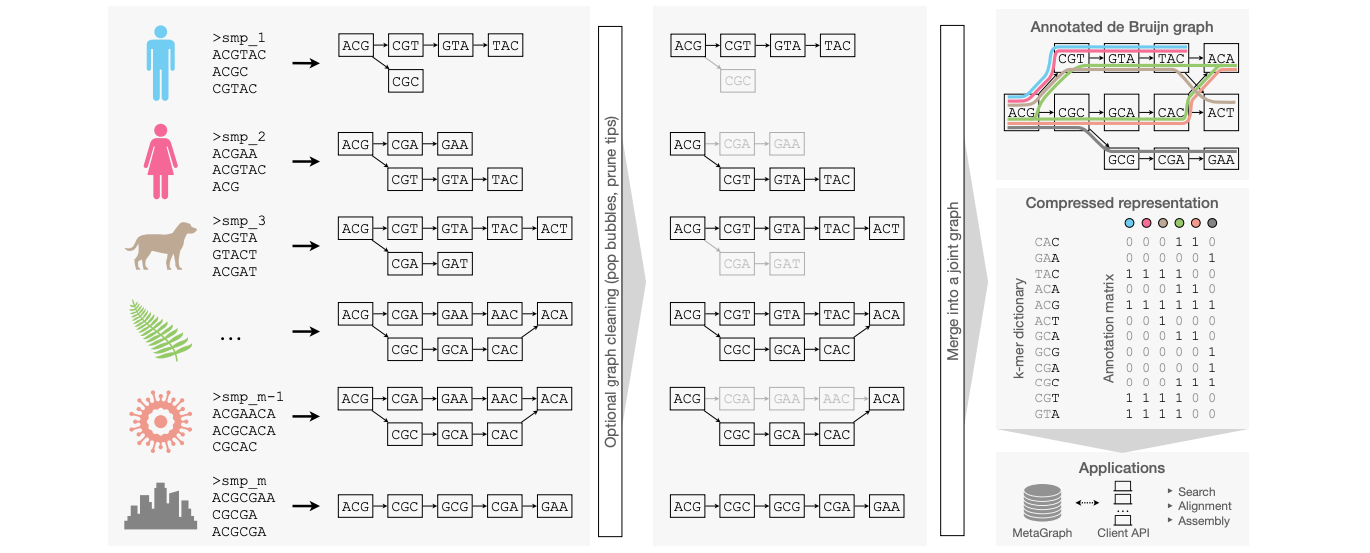
Indexing All Life's Known Biological Sequences
There's a lot of biological data out there but how can you search it? This paper suggests a methodology using annotated De Bruijn graphs to scalably index very large sets of DNA or protein sequences. They use efficient data structures and algorithms to compress Petabases of DNA sequences and make the indexes available for the community and show that these indexes can be useful for different kinds of analyses.
Open Source
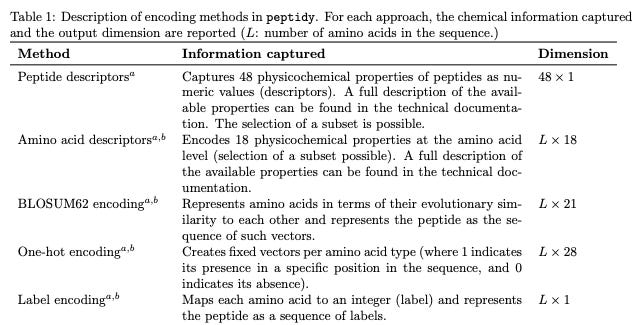
Peptidy: A Light-Weight Python Library for Peptide Representation in Machine Learning
This work presents a Python library to convert peptides (including those with post-translational modifications) into numerical representations that can be fed into ML models. The library is lightweight, has no external dependencies, and supports a range of encoding strategies.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
CARE
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋