Portal Weekly #51: ML for drug discovery summer school, using BioNeMo to fold the human proteome, TrogoTACs, and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
📅 Events Recap
ML for Drug Discovery Summer School
Last week, we kicked off the inaugural ML for Drug Discovery Summer School in Montreal. 150+ participants from across the world were in attendance to go through 5 days of programmed lectures:
Day 1 - Foundations and ML in Ligand-Based Modeling
Day 2 - ML in Structure-Based Drug Discovery
Day 3 - Generative Models and Molecular Design
Day 4 - Target Discovery and Deconvolution
Day 5 - Frontiers in AI for Drug Discovery
See the full schedule here. Looking to access the lecture slides and recordings? DM Jonny on Portal.
We’re currently in the middle of the summer school hackathon. Students have formed teams of 5 to work on kinase selectivity and ADME property prediction tasks. Datasets and benchmarks are accessible on Polaris. Check-in later today to see which teams won!
2024 Molecular Machine Learning Conference
On Wednesday, we also hosted MoML 2024 at Mila. We had nearly 200 attendees and ~50 poster submissions. It was amazing to see the interest and excitement for TechBio. We had speakers like Jian Tang and Max Jaderberg from Isomorphic Labs there to discuss topics related to geometric deep learning for proteins to molecular property prediction to AlphaFold 3 and more. We’ll be making the recordings available soon on YouTube.
💻 Latest Blogs
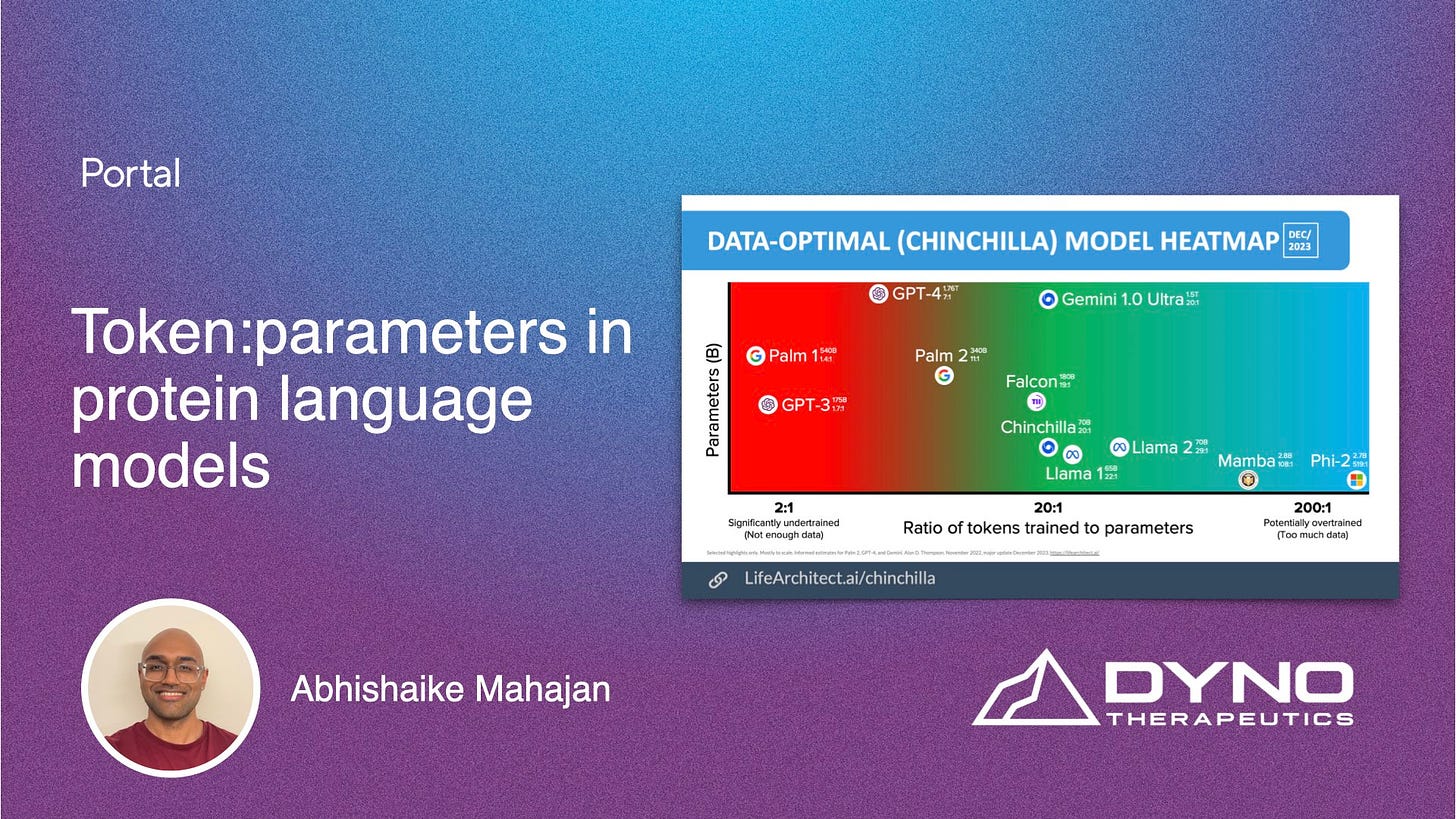
In the latest blog, Abhishaike Mahajan takes a brief look at scaling in current protein language models, specifically at the relationship between tokens trained and parameter count. Check it out here.
💬 Upcoming talks
LoGG continues next week with a talk by Timothy Duignan, who will demonstrate that Neural Network Potentials can reliably be recursively trained on a subset of their own output to enable coarse-grained continuum solvent molecular simulations that can access much longer timescales.
Join us live on Zoom on Monday, June 24th at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter and LinkedIn!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
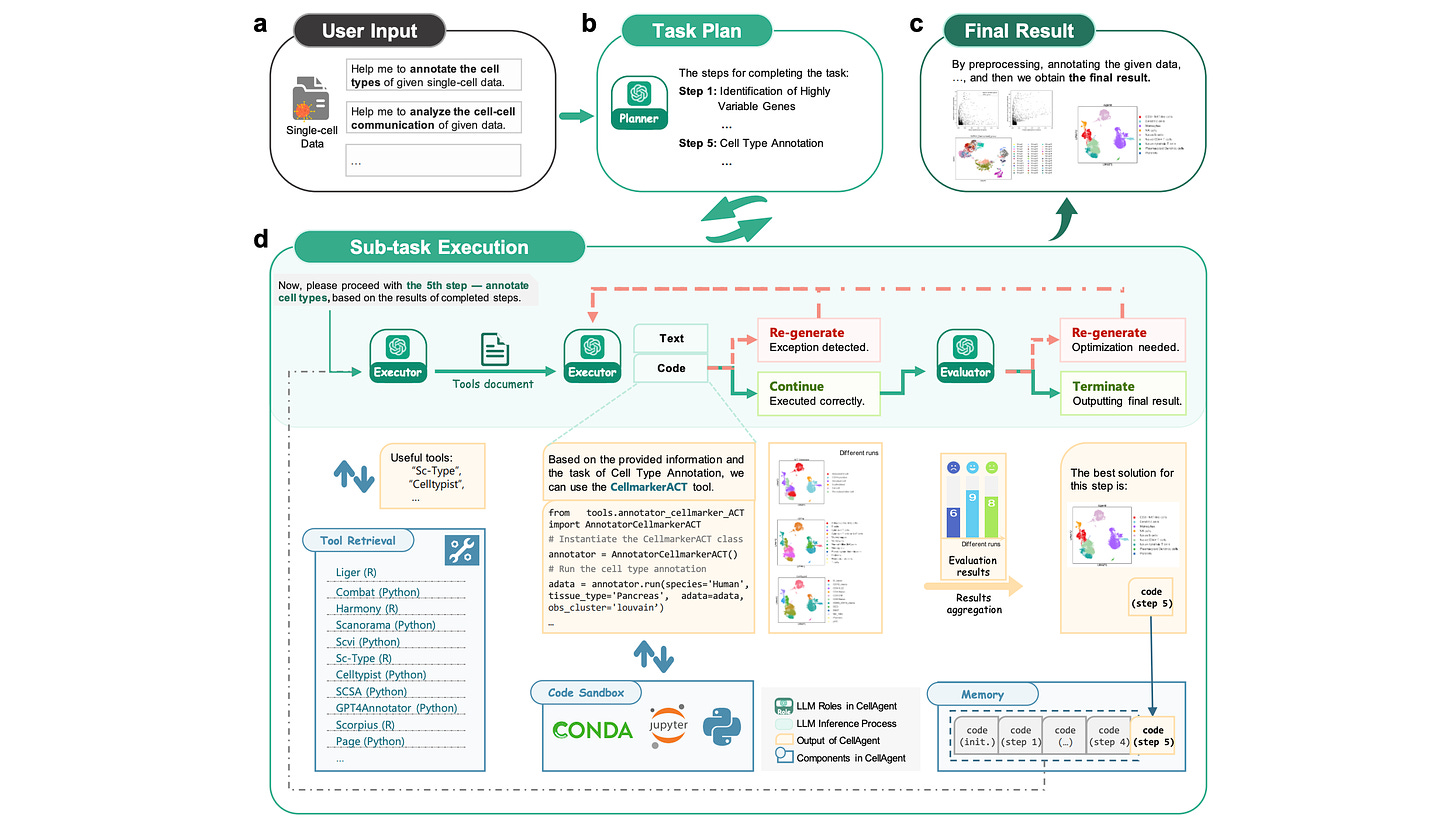
CellAgent: An LLM-driven Multi-Agent Framework for Automated Single-Cell Data Analysis
Stringing together tools for a single-cell analysis pipeline can be laborious. This study proposes an LLM-driven multi-agent framework specifically designed for this task. Different ‘biological expert roles’ are created and coordinated , with a self-iterative optimization mechanism allowing the agent to autonomously work on solutions. Their results suggest that CellAgent consistently identifies suitable tools and hyperparameters, providing another step towards self-driving laboratories.
Interested in self-driving labs and their potential? Check out this recent M2D2 talk by Alán Aspuru-Guzik!
ML for Small Molecules
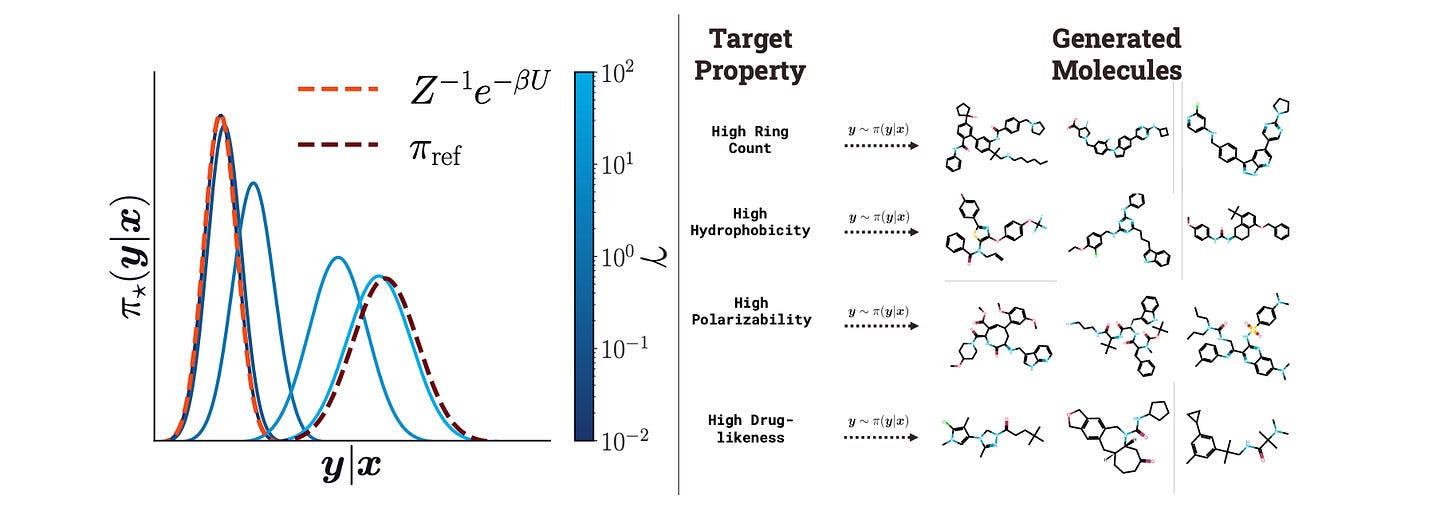
Energy Rank Alignment: Using Preference Optimization to Search Chemical Space at Scale
Generating molecules is hard in part because of the massive chemical search space. This study introduces energy rank alignment (ERA), a scalable algorithm that doesn’t use reinforcement learning and can be used to optimize autoregressive policies. Interestingly, the techniques described here, although designed for chemical search, seem to do well on an AI supervised task for LLM alignment, suggesting that the approach is quite generalizable.
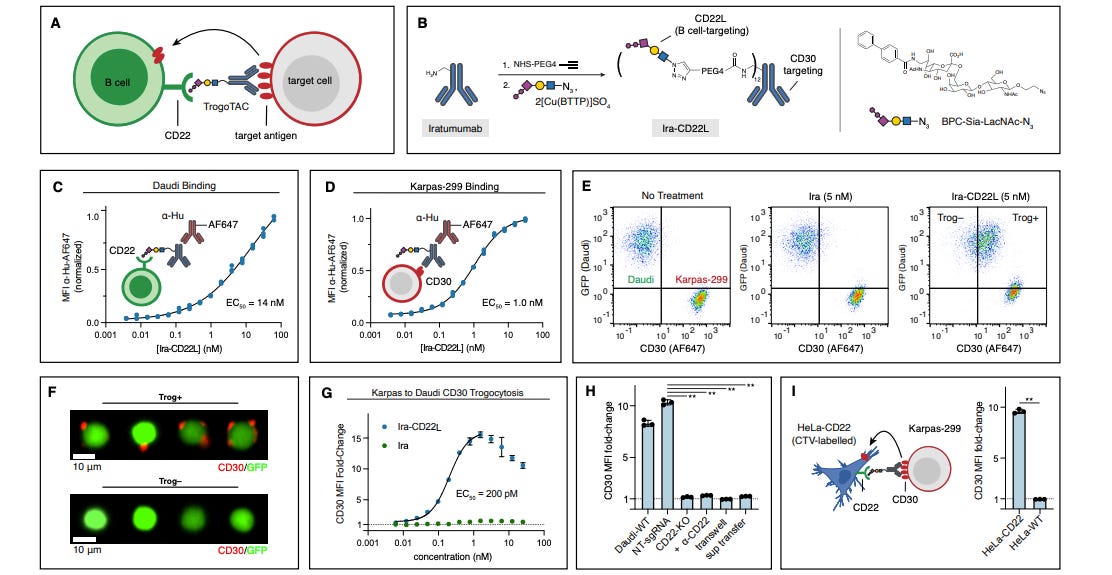
Trogocytosis Targeting Chimeras (TrogoTACs) for Targeted Protein Transfer
Trogocytosis is a type of cell-cell interaction that happens particularly with immune cells, where one cell contacts and ‘nibbles’ another cell; this transfers membrane fragments from one to the other. This study’s authors wondered if they could take advantage of this phenomenon to restore cells deficient in cell-surface proteins. They report the development of TrogoTACs, molecules that seem to be able to induce protein transfer between distinct cells. This could open up new therapeutic avenues, especially for cases where genetic intervention is hard.
ML for Atomistic Simulations
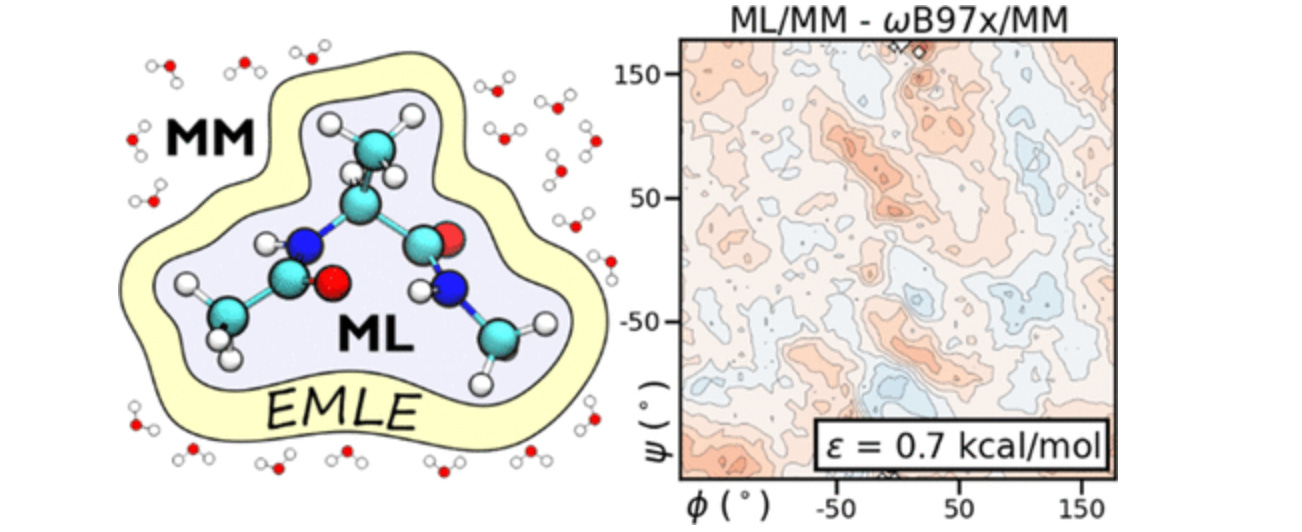
emle-engine: A Flexible Electrostatic Machine Learning Embedding Package for Multiscale Molecular Dynamics Simulations
This study presents a new embedding scheme for hybrid machine learning potential/molecular-mechanics (ML/MM) dynamics simulations, with systematic reductions in average error of the free energy surface when compared to MM embedding. They provide a package that can be used in existing QM/MM software, enabling current workflows to take advantage of these advances.
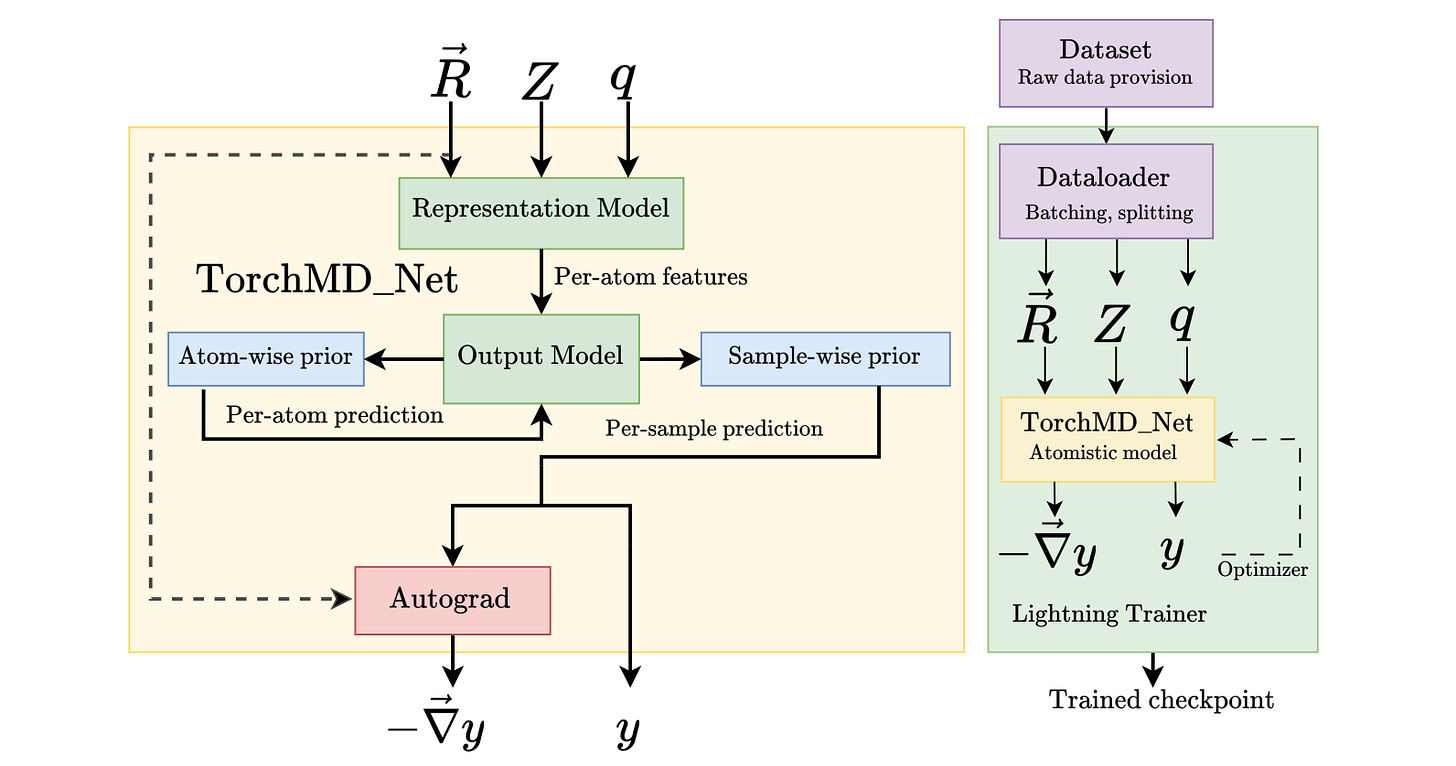
TorchMD-Net 2.0: Fast Neural Network Potentials for Molecular Simulations
This study presents advancements in the TorchMD-Net software package, incorporating cutting-edge architectures through a modular design approach and increasing computational efficiency. They have also included optimized neighbor search algorithms that support periodic boundary conditions and integration with existing molecular dynamics frameworks. The updated version can also integrate physical priors, which intuitively should give benefits in different areas of research.
ML for Proteins
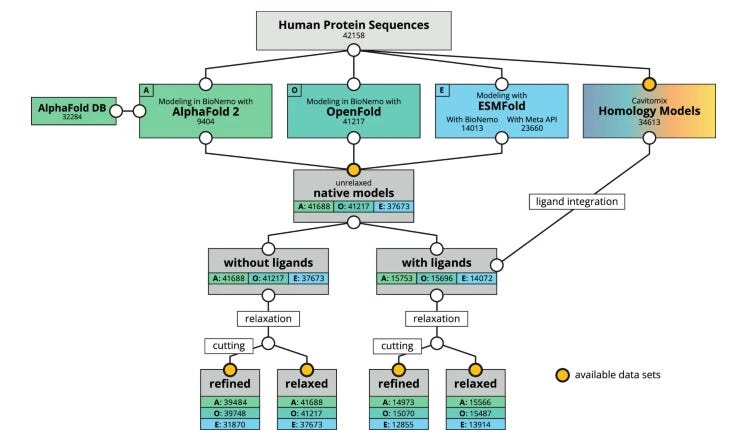
Folding the Human Proteome Using BioNeMo: A Fused Dataset of Structural Models for Machine Learning Purposes
NVIDIA’s BioNeMo platform provides state-of-the-art biomolecular models for AI drug discovery and a training service with data loaders and training workflows. This study combines modeling with AlphaFold 2, OpenFold, ESMFold and Innophore’s CavitomiX platform to generate a dataset of predicted protein structures for 42,042 distinct human proteins, including splicing variants, for a reference proteome. They provide both edited and unedited formats, giving a potentially powerful dataset for different research purposes.
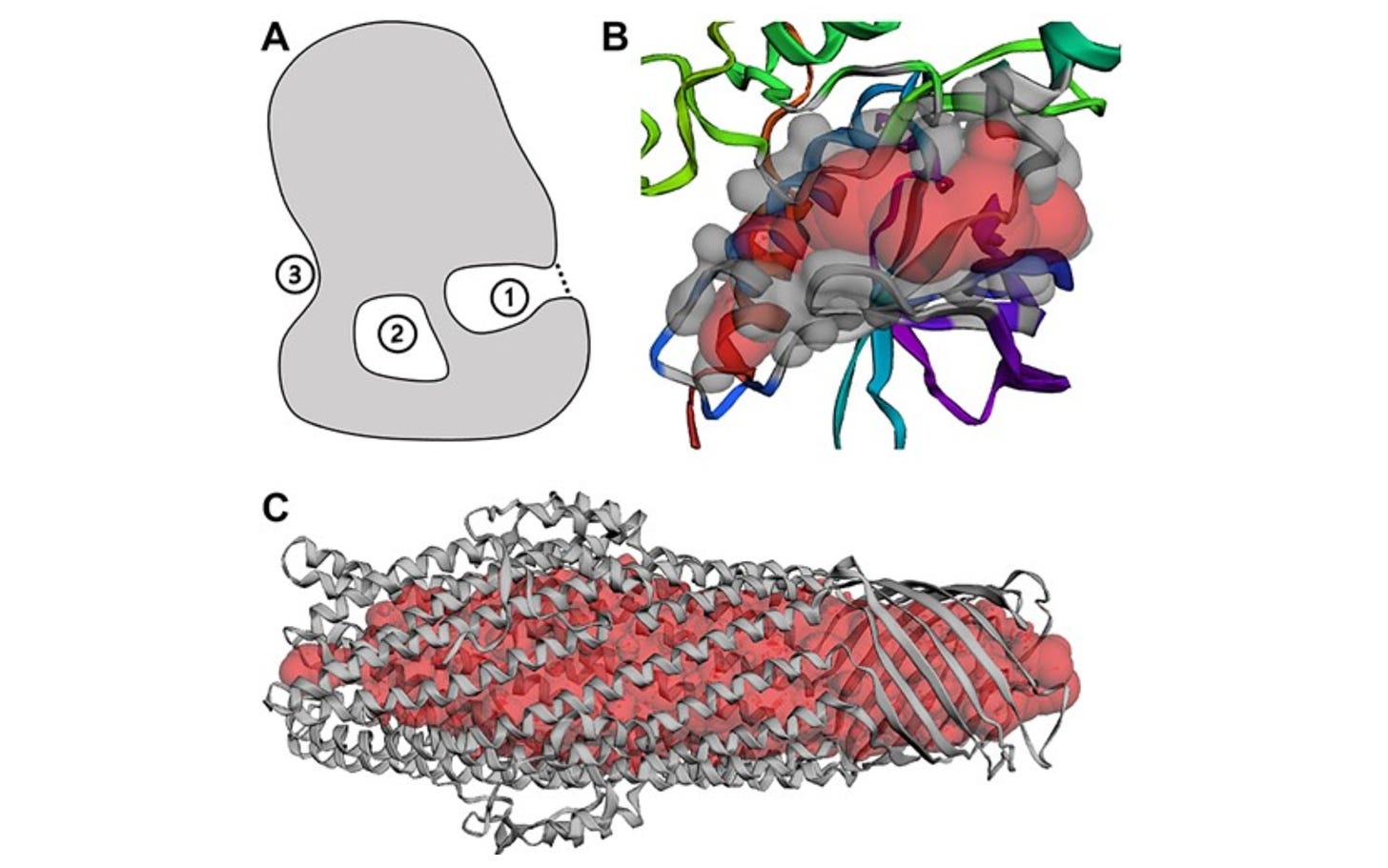
CASTpFold: Computed Atlas of Surface Topography of the Universe of Protein Folds
CASTp is a widely used web server for locating, delineating, and measuring geometric and topological properties of protein structures. This extension gives results on an expanded database of proteins, including the Protein Data Bank (PDB) and 183 million AlphaFold2 structures. It also gives functional pockets prediction with Gene Ontology terms, and a pocket similarity search function.
ML for Omics
GenBench: A Benchmarking Suite for Systematic Evaluation of Genomic Foundation Models
Genomics is hard, and complicated. The genomics field still seems to be settling on benchmarks to properly evaluate proposed foundation models in this space. This study presents one such benchmarking attempt, presenting a set of tasks broken up as ‘short range’ or ‘long range’ and evaluating a number of current high-profile genomic foundation models like Caduceus, DNABERT, HyenaDNA and Nucleotide Transformer, alongside some more ‘expert’ models.
The tasks presented in this study are more numerous but less varied than BEND - check out Frederikke and Felix’s blog post for more details on that work!
Open Source
BioMoDes: A Repository of Tools for Biomolecular Modeling and Design
This site provides an overview of different tools for structure prediction, design, protein search and more, mainly for tools published in 2024.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
Contextual AI Models for Single-cell Protein Biology by Michelle Li
gRNAde: Geometric Deep Learning for 3D RNA inverse design by Chaitanya K. Joshi
LoGG
Approximately Piecewise E(3) Equivariant Point Networks by Matan Atzmon
CARE
Identifying Representations for Intervention Extrapolation by Sorawit (James) Saengkyongam
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋