Portal Weekly #52: ESM3 generative protein language model, Therapeutics Data Commons goes multimodal, computational venom design and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
💻 Latest Blogs
In the latest blog, Song Xia and Eric Chen give an insight into tools being developed in Yingkai Zhang’s lab at NYU, for pocket-guided rational design approaches, protein−ligand docking and more.
💬 Upcoming talks
LoGG continues next week with a talk by Kacper Kapusniak, who will present a framework for getting the generative model to match vector fields on the data manifold, giving lower uncertainty and more meaningful interpolations.
Join us live on Zoom on Monday, July 1st at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter and LinkedIn!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
Using Large Language Models for Safety-Related Table Summarization in Clinical Study Reports
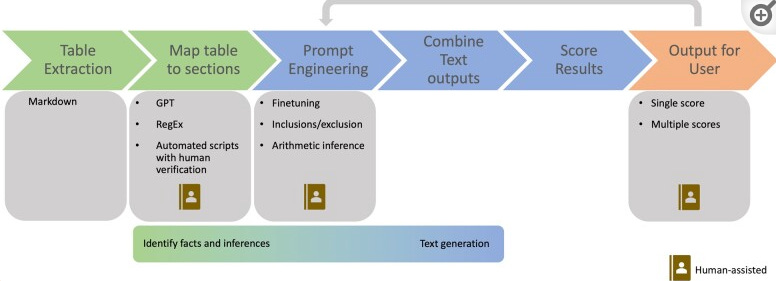
Clinical study reports (CSRs) are often created as part of the process of submitting applications for new medical treatments to regulators. CSRs answer questions such as: Why was the trial done? What were the important questions asked in the trial? What were the results? As part of a challenge initiated by Pfizer, several teams created a pilot for generating summaries of safety tables for clinical study reports (CSRs). The challenge results show the potential of LLMs in automating table summarization in CSRs, but also the importance of human involvement and further research.
ML for Small Molecules
Molecular Display of the Animal Meta-Venome for Discovery of Novel Therapeutic Peptides
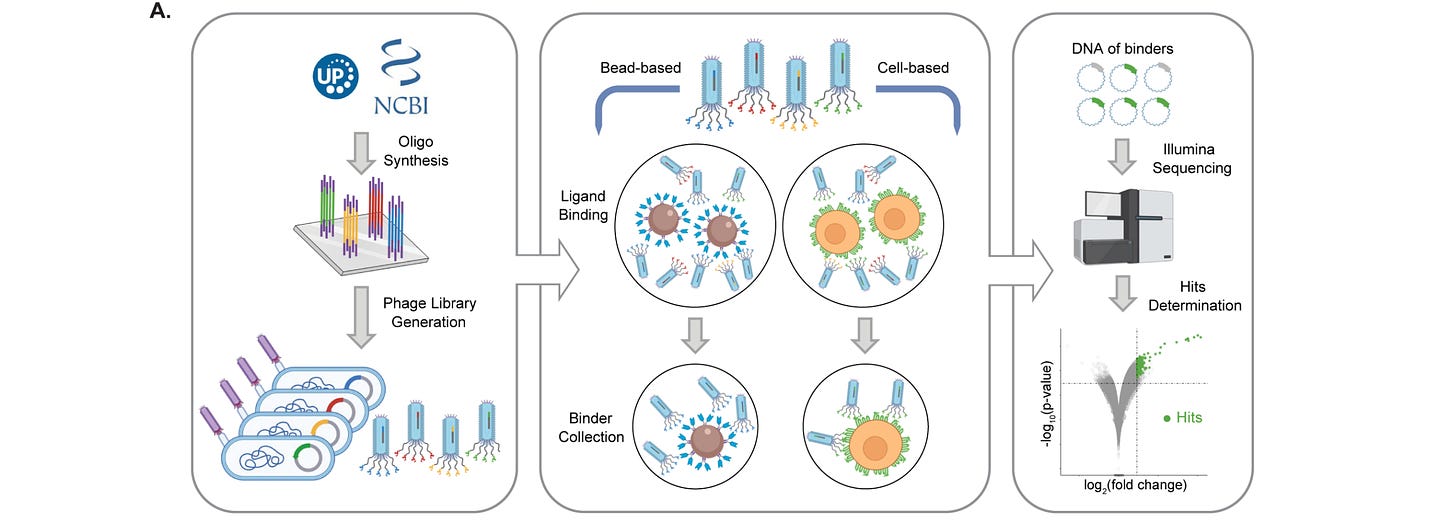
Animal venoms have evolved to have specific and strong effects, but have not really been considered as therapeutics. This is partially because it’s hard to get enough venoms/venom-like molecules to evaluate by high-throughput screening. This study computationally designed a diverse library of animal venoms and ‘metavenoms’, discovering new proteins that target specific receptors (in their case, an ‘itch receptor’) and also used deep learning to identify two natural human versions that affect their ‘itch receptor’, paving the way for a more high-throughput screening of animal venoms for therapeutic purposes.
ML for Atomistic Simulations
Automated Adaptive Absolute Binding Free Energy Calculations
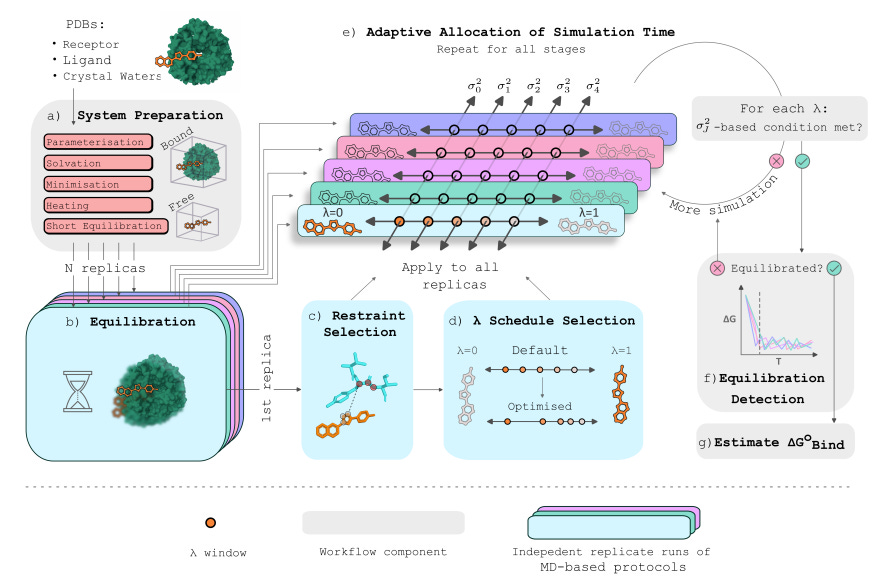
Removing the need for human intervention in workflows can speed up research, as long as the automated workflow is robust enough. This study introduces an automated alchemical absolute binding free energy workflow that selects windows, detects equilibration using an ensemble, and allocates sampling time based on inter-replicate statistics. They believe that they have chosen reasonable default parameters and that the automated pipeline is simple to implement, and have provided an open-source package to help in using the tool.
ML for Proteins
Simulating 500 million years of evolution with a language model
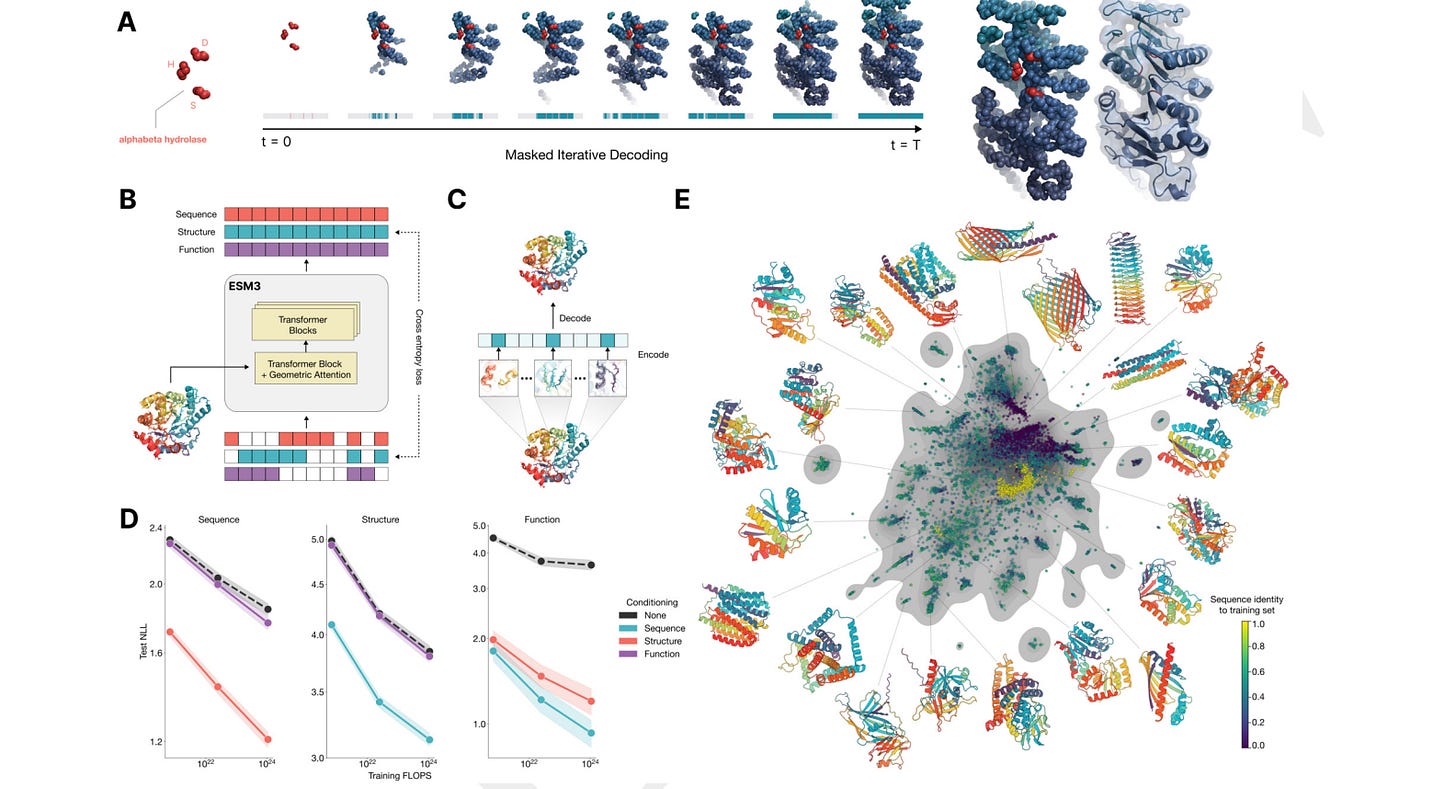
ESM-2 (Evolutionary Scale Modeling) from MetaAI is an impactful protein language model, up there with AlphaFold 2. The latest iteration, ESM3, as announced in a blog post, was trained on almost 2.8 billion protein sequences. A small open version for non-commercial use is coming soon to NVIDIA’s BioNeMo platform and is available through HuggingFace, according to the ESM3 GitHub repo. You can apply for beta access to the full family of ESM3 models at EvolutionaryScale Forge.
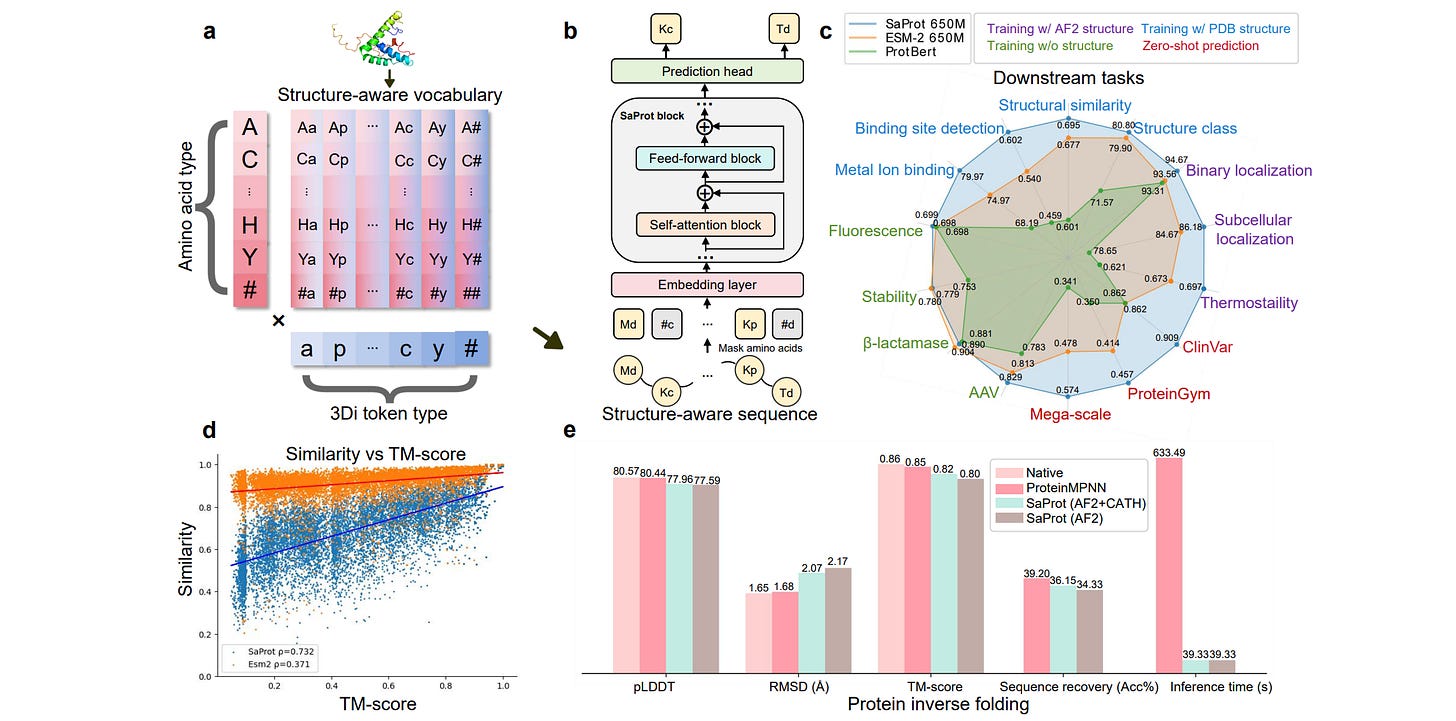
SaprotHub: Making Protein Modeling Accessible to All Biologists
Building on their SaProt work, Jin Su and colleagues have now put out a platform for training, using and sharing protein language models. The Hub consists of the Saprot protein language model, ColabSaprot, and an open community store for saving and sharing Saprot models. It’s an initiative to promote more collaboration and take steps towards an “Open Protein Modeling Consortium”.
Jin Su wrote a blog post here, if you haven't seen it!
ML for Omics
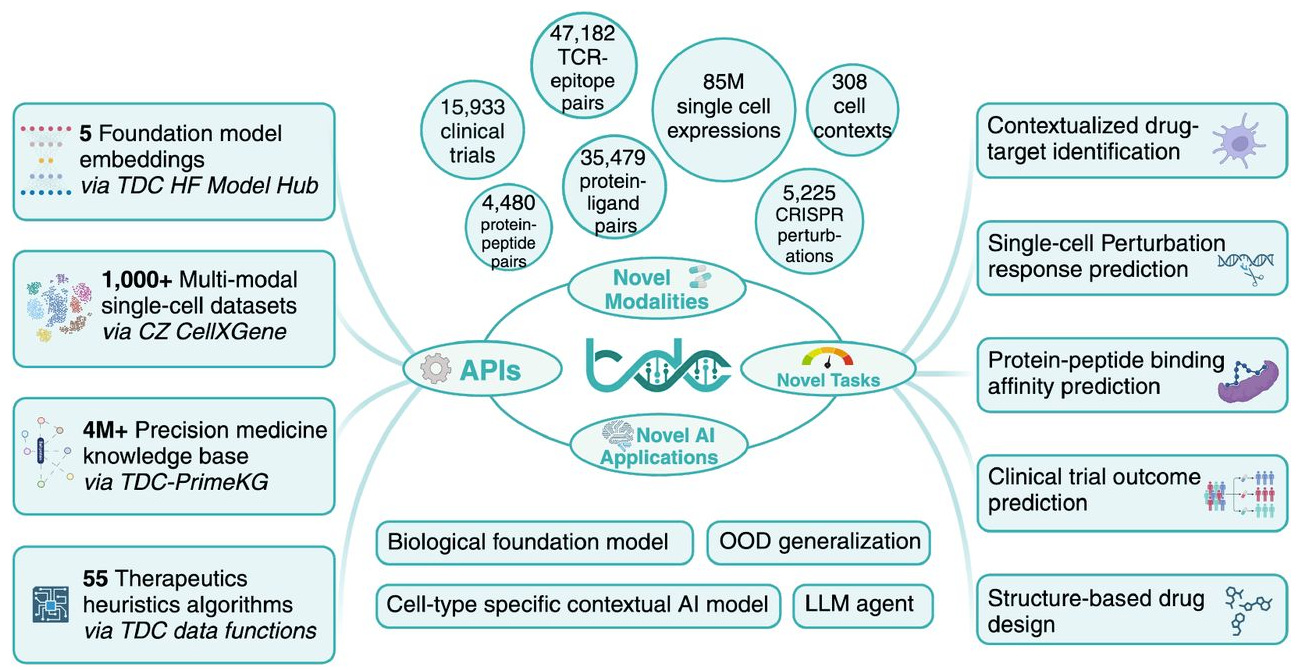
TDC-2: Multimodal Foundation for Therapeutic Science
The Therapeutics Data Commons is already known a hub for datasets for drug discovery, with leaderboards encouraging competition in ADMET prediction and other tasks. This preprint describes an expansion (or rather, an ‘overhaul’) of TDC to include >1,000 multimodal datasets comprising 85 million cells, pre-calculated embeddings from five single-cell foundation models, and a biomedical knowledge graph. In line with this expansion, there are seven new ML tasks, including contextualized drug-target identification, single-cell chemical/genetic perturbation response prediction, and clinical trial outcome prediction. They also release new benchmarks evaluating state-of-the-art models; altogether, this should be a good resource for those looking to test models for different purposes.
Models trained on TDC, like ADMET-AI, perform well in a range of tasks. Check out Kyle Swanson’s blog post for more insight into ADMET-AI!
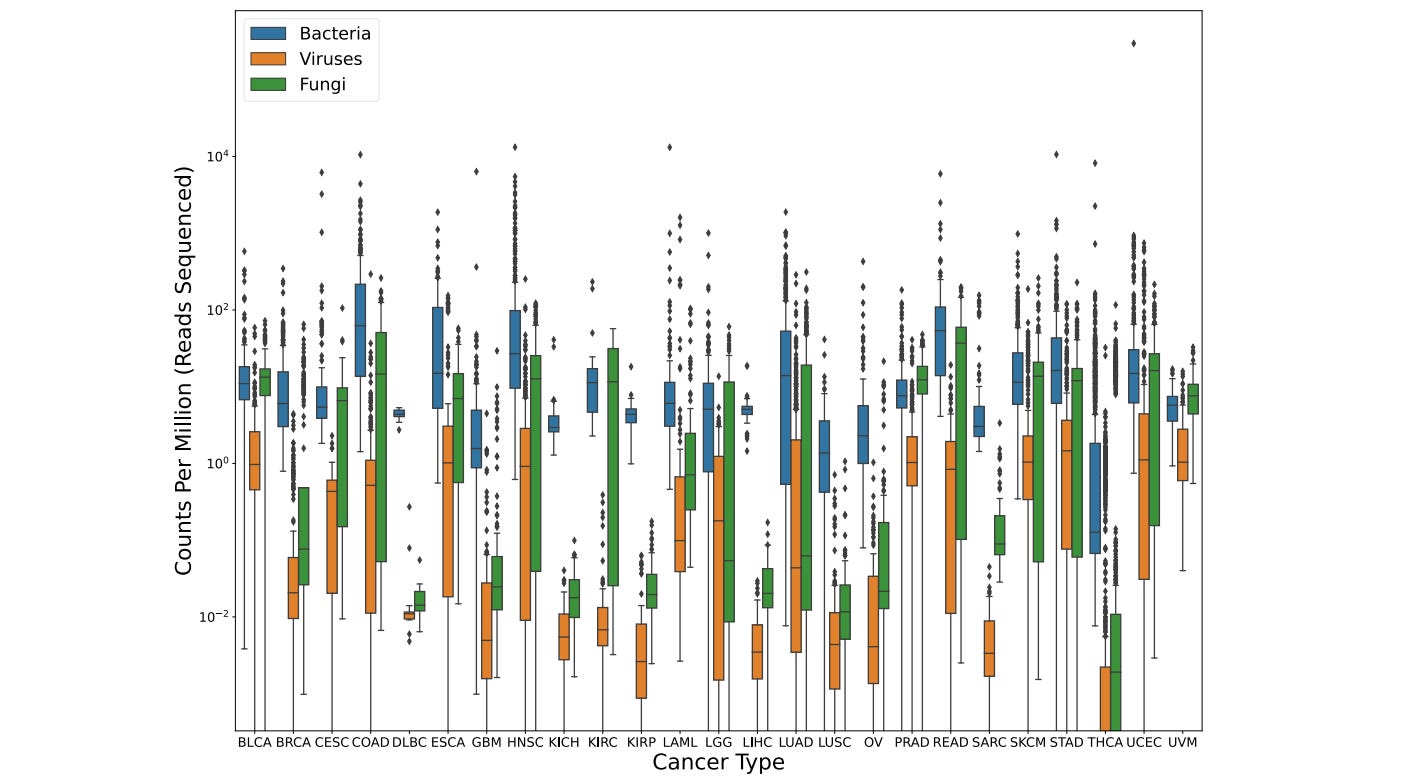
Comprehensive Analysis of Microbial Content in Whole-Genome Sequencing Samples from The Cancer Genome Atlas Project
Many recent publications have reported the presence of microbial species in human tumors. This study suggests that those reports were overblown, and the presence of microbes is far smaller; many are known contaminant. The authors release a dataset containing detailed read counts for bacteria, viruses, archaea, and fungi detected in all ~5700 samples from The Cancer Genome Atlas, which they hope will serve as a public reference for the future.
Open Source
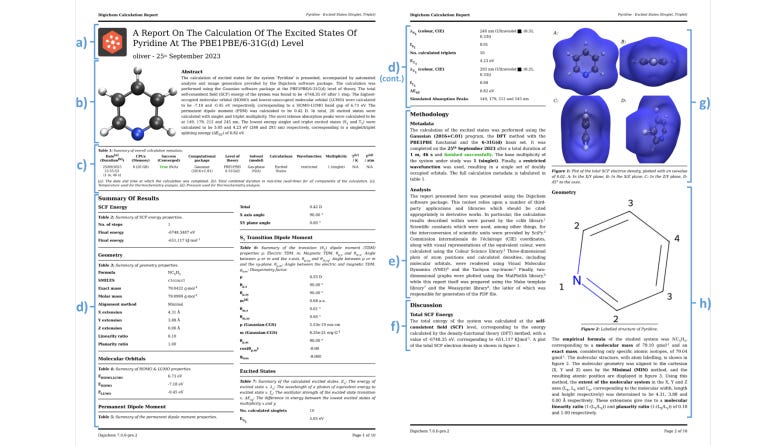
Digichem: Computational Chemistry For Everyone
To avoid human error, we should try to automate as many pipelines as we can. This preprint introduces a program that tries to simplify and automate the computational chemistry pipeline including generation of 3D density plots and 2D graphs of calculation data.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
LoGG
Scalable molecular simulation of electrolyte solutions with quantum chemical accuracy by Timothy Duignan
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋