🪗 Community Round-Up: Week of October 16th, 2023
M2D2 Talks are BACK! Also: benchmarks, EVEscape, self-improving LLMs and many more papers
Hi everyone 👋
Welcome to another issue of the community newsletter! With so much happening in the field of AI for drug discovery, we’re now increasing the cadence of the newsletter to once a week. For better organization, we’ve done our best to categorize papers into key topics: ML for Small Molecules, ML for Atomistic Simulations, Multi-Modal Learning, ML for Proteins, LLMs for Science, and Open Source.
We’re also excited to announce that the M2D2 Talks are back! 🚀 We’ll be joined by Hannes Stärk from MIT who will discuss his recent work on flow models for multi-ligand docking and binding site design.
Join us live on Zoom on Tuesday, October 24th at 11 am ET. Find more details here.
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
ML for Small Molecules
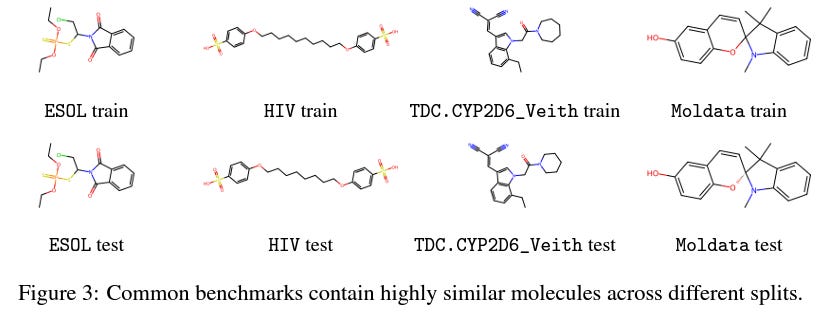
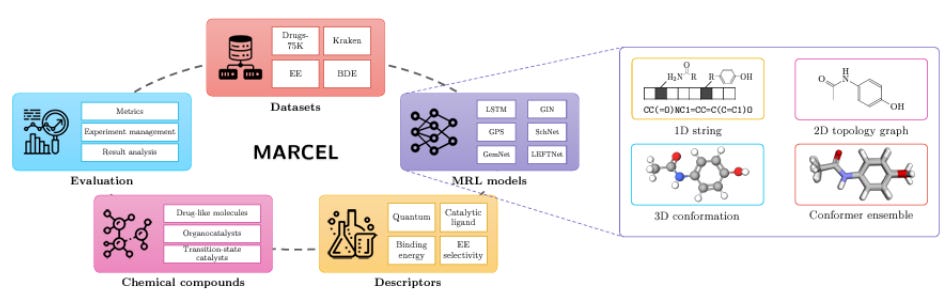
There is a lot of discussion in the AI for Drug Discovery community about benchmarks: the general feeling is that existing benchmarks are lacking in some ways and we need more real-world evaluations for our models. Among recent work addressing this issue, Lo-Hi: Practical ML Drug Discovery Benchmark proposes a benchmark for “the real drug discovery process” tasks of Lead Optimization and Hit Identification; Learning Over Molecular Conformer Ensembles: Datasets and Benchmarks proposes a benchmark to evaluate models that try to account for molecular flexibility by learning on conformer ensembles, and Genetic Algorithms are Strong Baselines for Molecule Generation argues that genetic algorithms can outperform more complicated ML models in molecule generation, so new algorithms should have some clear advantage over them.
Figure 3 from the Lo-Hi paper
Figure 1 from the Molecular Conformer paper
ML for Atomistic Simulations
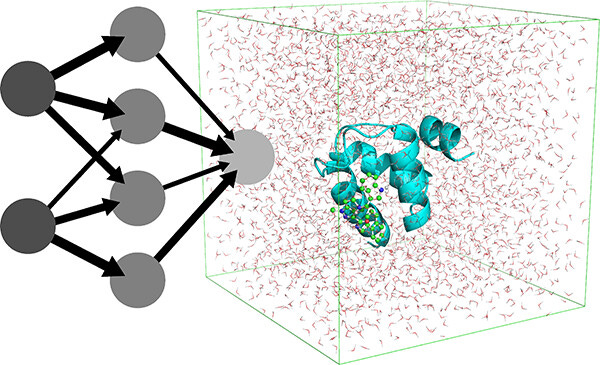
NNP/MM: Accelerating Molecular Dynamics Simulations with Machine Learning Potentials and Molecular Mechanics
ML potentials can improve the accuracy of biomolecular simulations but are limited by computational costs. This study introduces an optimization of the hybrid method that combines a neural network potential with molecular mechanics. They report ~5x increased simulation speeds and longer simulations, which are promising improvements for this class of simulations.
EGraFFBench: Evaluation of Equivariant Graph Neural Network Force Fields for Atomistic Simulations
Equivariant graph neural networks force fields (EGraFFs) exploit graph symmetries for modelling complex interactions in atomic systems. There are now a lot of EGraFF architectures out there, but fewer studies comparing these models on real-world atomistic simulations. This study systematically benchmarks 6 EGraFF algorithms and also releases two new benchmark datasets, establishing a framework for evaluating ML force fields and pointing to open research challenges in this area.
Multi-Modal Learning
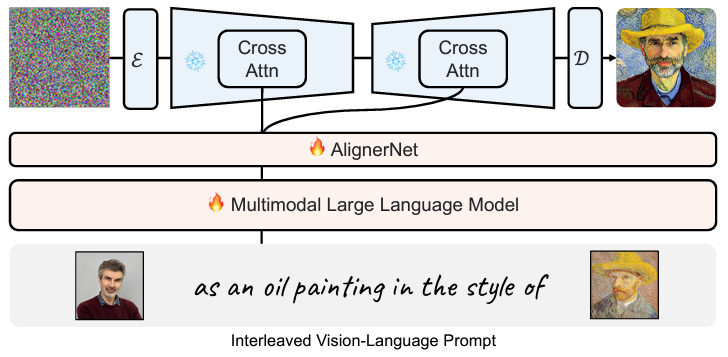
KOSMOS-G: Generating Images in Context with Multimodal Large Language Models
Text-to-image and vision-language-to-image generative models have progressed markedly, but generation from vision-language inputs, especially involving multiple images, is under-explored. This study proposes KOSMOS-G, which leverages multimodal LLMs and CLIP using text as an anchor, resulting in better zero-shot multi-entity subject-driven generation.
ML for Proteins
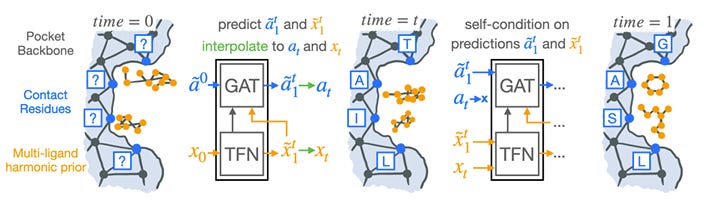
Harmonic Self-Conditioned Flow Matching for Multi-Ligand Docking and Binding Site Design
Many proteins are activated by small molecules, or act on them for catalysis. Designing binding pockets consequently has a range of potential applications. This study presents two generative flow models that improve on state-of-the-art generative processes and provide the first general solution for binding site design.
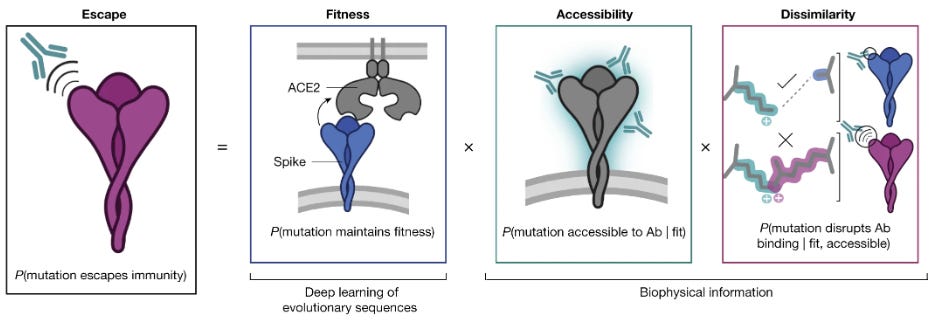
How AlphaFold And Other AI tools Could Help Us Prepare For The Next Pandemic
Modeling the structure of a protein, both in its natural form and with mutations, can help us to understand how that protein works. This Nature News article highlights a number of ways our response to viruses can be ameliorated with ML, and spotlights EVEscape, a modelling framework combining deep learning models and biophysical constraints trained on coronavirus sequences from before January 2020 that is remarkably successful in predicting the evolution trajectory of real-world SARS-CoV-2 variants past that date.
LLMs for Science
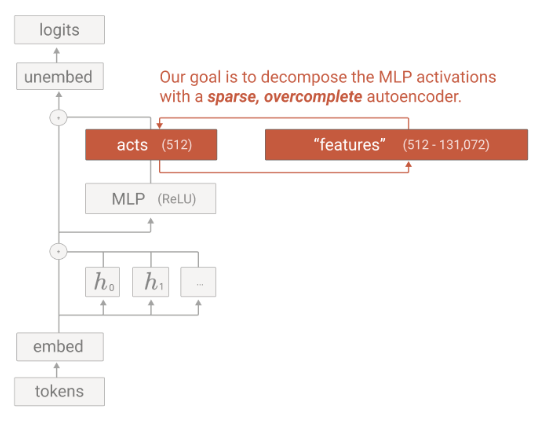
Towards Monosemanticity: Decomposing Language Models With Dictionary Learning
Explainability is an important part of modern ML. By understanding what individual components are doing and how they interact, we can better understand the whole network. This study uses a sparse autoencoder to extract interpretable features and enable circuit analysis. They suggest that this approach can be used to intervene on and steer transformer generation, and find features that would otherwise not appear using a neuron-centric approach.
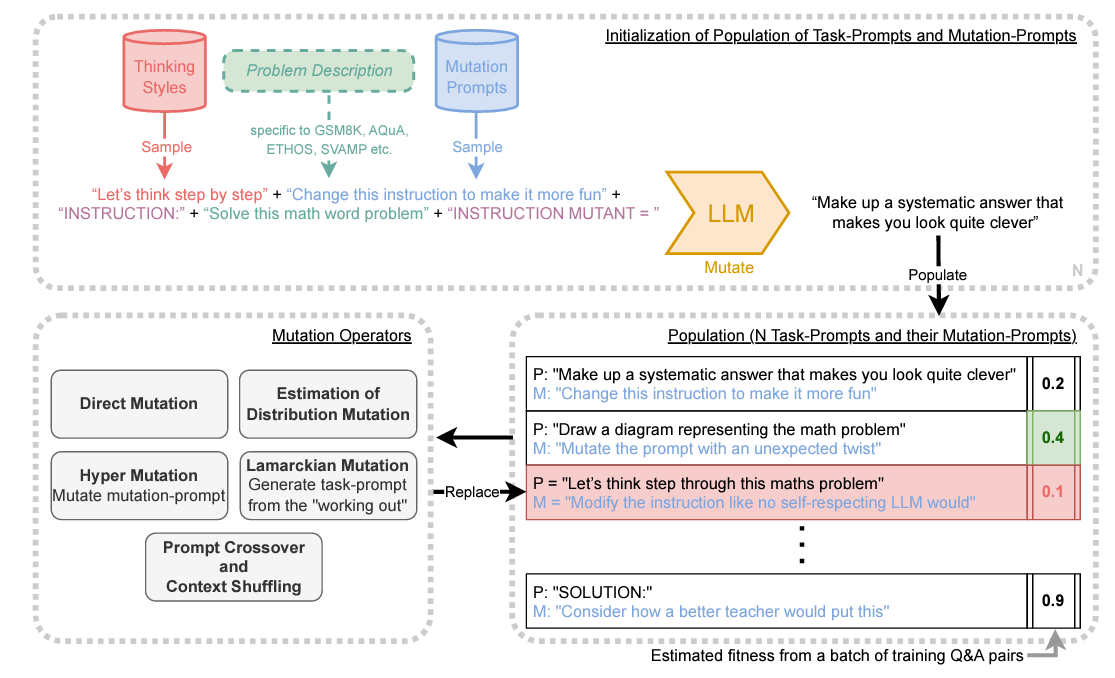
Promptbreeder: Self-Referential Self-Improvement via Prompt Evolution
This paper discusses PromptBreeder, a novel self-referential self-improvement mechanism for enhancing prompts in Large Language Models (LLMs). Unlike hand-crafted prompt-strategies, PromptBreeder evolves and adapts prompts for specific domains using LLM guidance. It mutates a population of task-prompts, evaluates them for fitness, and iterates over generations to evolve them. Remarkably, PromptBreeder also improves the mutation-prompts that enhance task-prompts, outperforming established prompt strategies on arithmetic and commonsense reasoning benchmarks.
ML for Omics
A Collection of Single-Cell Foundation Models
Foundation models are large machine learning models trained on a vast quantity of data, with the ability to be adapted to a range of downstream tasks. This GitHub repository aims to bring together different foundation models for single-cell biology, with the idea that a successful foundational cell model will integrate multiple modalities and be able to predict unseen, new cell states across health and disease.
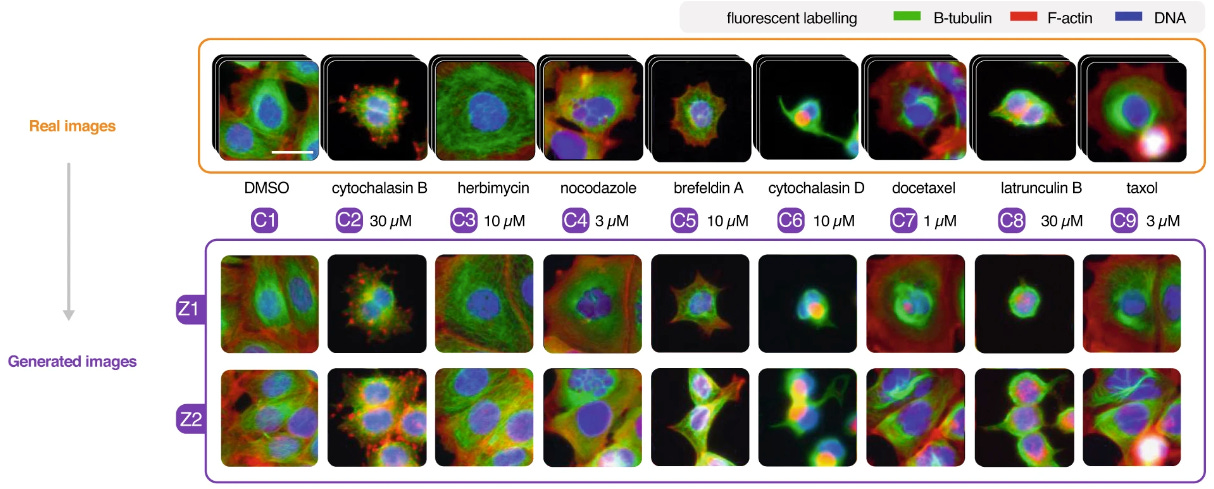
Revealing Invisible Cell Phenotypes with Conditional Generative Modeling
Biological experiments rely on performing a perturbation and observing some kind of effect, often via microscopy for cell studies. However, it is often hard to spot the effects due to heterogeneity and effect size. This study uses conditional generative models to transform an images of cells in one condition to another, canceling cell variability. This approach could make biological and disease markers easier to discover.
Open Source
Kaggle competition - Single-Cell Perturbations
Kaggle competitions can be a good way to try implementing some of the techniques covered in our newsletters, or to explore the space. This challenge, part of the NeurIPS2023 Competition Track, is to predict expression from an incomplete cell-type x perturbation matrix.
Think we missed something? Join our community to discuss these topics further!
Valence Portal is the home of the AI for drug discovery community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋