Portal Weekly #25: NeurIPS event with Recursion, NVIDIA, and Valence Labs, biomedical language models, multi-omics Mowgli, and more.
Join Valence Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on everything happening in TechBio research!
📅 Upcoming events
Recursion, NVIDIA, and Valence Labs are cohosting a 🎉TechBio social🎉 at NeurIPS this year! We want to extend an early invitation to our newsletter subscribers before we go live with public announcements next week. RSVP here to secure your spot - space is limited!
Join the event for an evening of drinks, bites, and great conversation on how AI/ML can be used to accelerate the drug discovery process.
💬 Upcoming talks
M2D2
M2D2 continues next week with a presentation by Martin Buttenschoen from Oxford University on scrutinizing claims of state-of-the-art performance in protein-ligand docking, and a way to address issues with implausibility.
Join us live on Zoom on Tuesday, November 28th at 11 am ET. Find more details here.
LoGG
LoGG continues with a presentation by Francisco Vargas from Cambridge University, who will talk about a framework for sampling and generative modelling centred around divergences on path space.
Join us live on Zoom on Monday, December 4th at 11 am ET. Find more details here.
CARE
We’re working on verifying details for next week’s speaker. Join the group live on Zoom on Wednesdays at 11 am ET. Check back here to see who the speaker will be!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
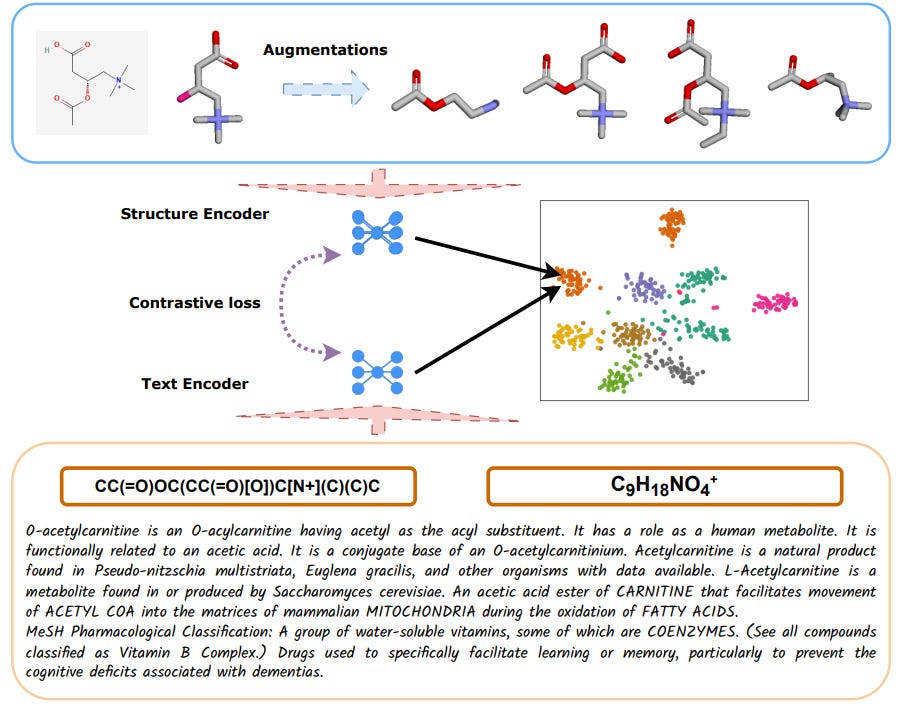
MolLM: Integrating 3D and 2D Molecular Representations with Biomedical Text via a Unified Pre-trained Language Model
Many people suspect that including 3D structural information will make deep learning models’ representations better, but most work relies on 1D or 2D formats. This study introduces a language model that incorporates biomedical text, 2D and 3D molecular information, using contrastive learning to learn across modalities. Not only do they show good representation capabilities across a number of downstream tasks, they also show through ablation that using 3D representations significantly improves performance.
MEDITRON-70B: Scaling Medical Pretraining for Large Language Models
Large language models (LLMs) could potentially improve access to medical knowledge, but current models are either closed-source or smaller-scale. This work introduces a suite of open-source LLMs, building on Llama-2 and pretrained on a curated medical corpus. The bigger MEDITRON outperforms GPT-3.5 and Med-PaLM and is approaching GPT-4 and Med-PaLM-2.
Graph Learning
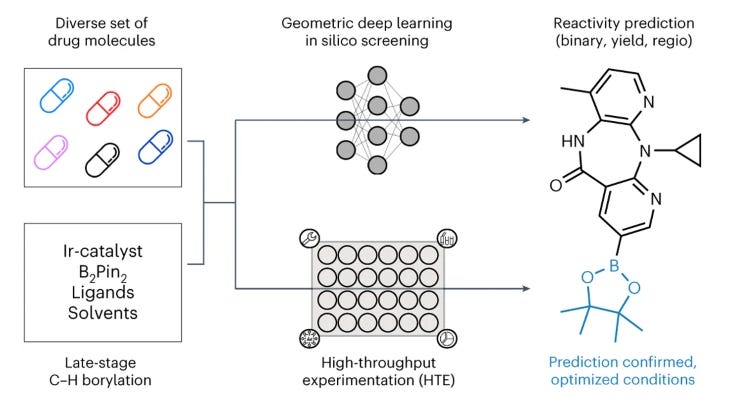
Enabling Late-Stage Drug Diversification by High-Throughput Experimentation with Geometric Deep Learning
In drug discovery, finding a potential drug and then improving its properties is expensive and time-consuming. Late-stage functionalization is one solution to this expense: during chemical synthesis, we want to transform a molecule without needing to add a functional group to enable that transformation. This study developed a platform using geometric deep learning and high-throughput screening that successfully identified opportunities for structural diversification.
ML for Small Molecules
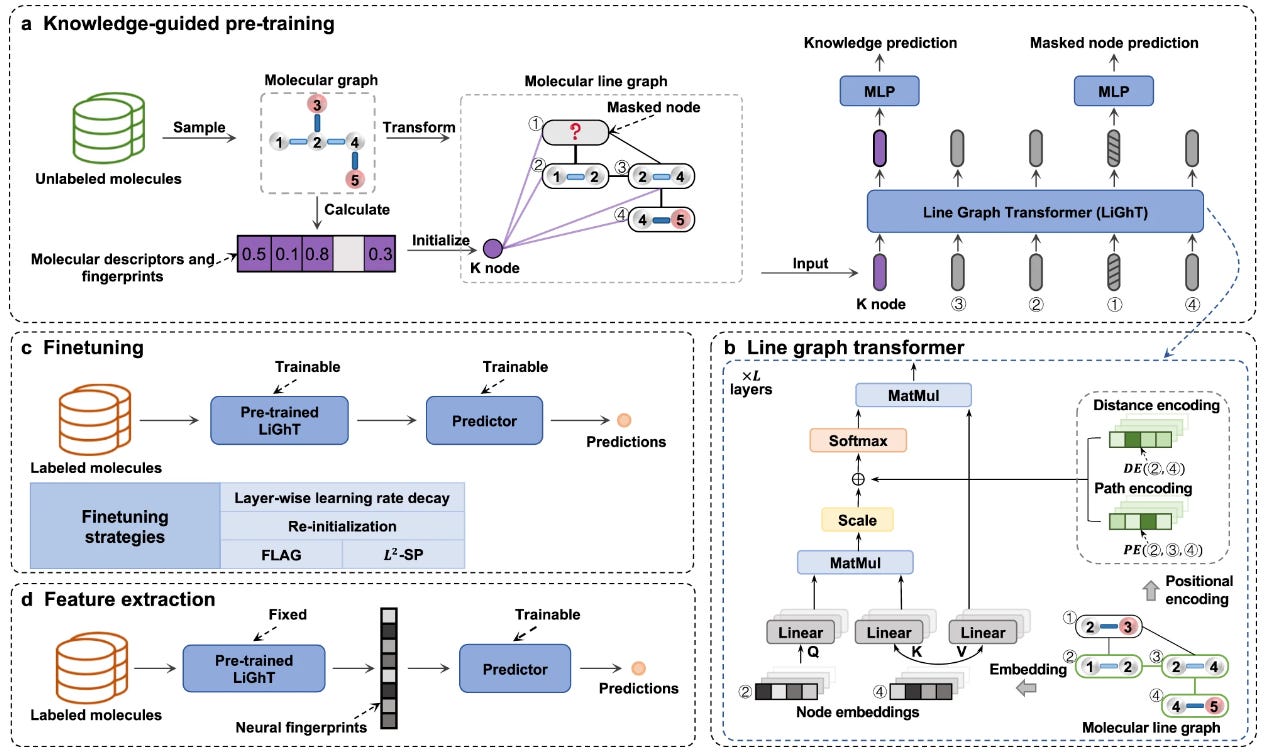
A Knowledge-Guided Pre-Training Framework for Improving Molecular Representation Learning
Self-supervised learning (SSL) is useful in the molecular domain because we often don’t have large datasets. Many strategies try to use SSL with graph neural networks (GNNs) for property prediction but are limited by the lack of a well-defined learning strategy and GNN capacity. This study combines a high-capacity line graph transformer with a knowledge node to try to capture structural and semantic information in molecular graphs. They show good performance on 63 molecular property datasets and identify potential inhibitors for two antitumor targets.
Chemical Complexity Challenge: Is Multi-Instance Machine Learning a Solution?
Molecules exist in 3D space and can have a bewildering variety of forms, and sometimes only one of those forms has bioactivity! How do we choose the right form for machine learning property prediction? This review suggests that multi-instance learning (MIL), where we represent an object as a set of alternative instances (a bag) and establish correlations between the bag of instances and the bag label. They describe MIL in the context of small molecules, and collect examples that show where MIL has an advantage over traditional single-instance learning.
ML for Atomistic Simulations
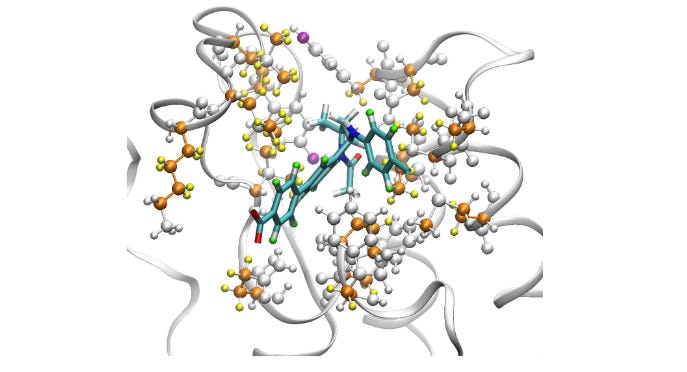
Global Ranking of the Sensitivity of Interaction Potential Contributions Within Classical Molecular Dynamics Force Fields
Uncertainty quantification (UQ) is being used in areas of computational science where we expect actionable outcomes. However, molecular dynamics (MD) doesn’t seem to use UQ as much. This study applies UQ to classical MD with a particular focus on uncertainties in the high-dimensional force-field parameters. They find that prediction uncertainty is dominated by a small number of the hundreds of interaction potential parameters.
ML for Proteins
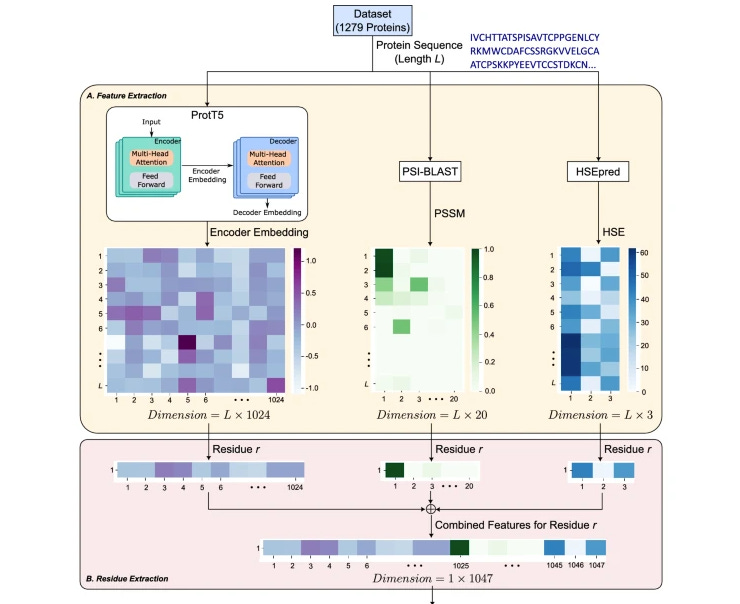
PepCNN Deep Learning Tool for Predicting Peptide Binding Residues in Proteins Using Sequence, Structural, and Language Model Features
Protein-peptide interactions play an important role in many cellular processes, and their dysfunction can lead to cancer and other problems. This study suggests PepCNN, a model that uses an embedding from a pre-trained protein language model combined with half-sphere exposure, a measure of the degree of surface exposure of an amino acid on a protein, and position-specific scoring matrices from multiple sequence alignments.
ML for Omics
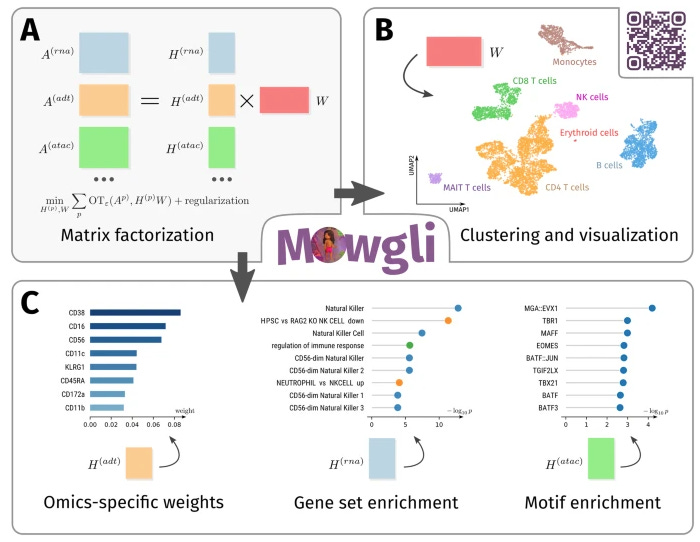
Paired Single-Cell Multi-Omics Data Integration with Mowgli
We can now profile multiple molecular layers from the same set of cells, but we need new methods to analyze them! This study introduces a method for integrating paired multi-omics data with any type and number of omics by combining integrative nonnegative matrix factorization and optimal transport. They also present an analysis that includes biological interpretability.
BEND: Benchmarking DNA Language Models on Biologically Meaningful Tasks
We have reams of genomic DNA sequences but annotating the different regions through experiments is laborious and challenging, so studies have tried to use DNA language modelling (DLM) instead. Evaluation of these models is often varied and unrealistic though. This study introduces a benchmark for DLMs that tries to be realistic and biologically meaningful. They find that DLMs can perform as well as experts on some tasks but struggle with long-range features.
Open Source
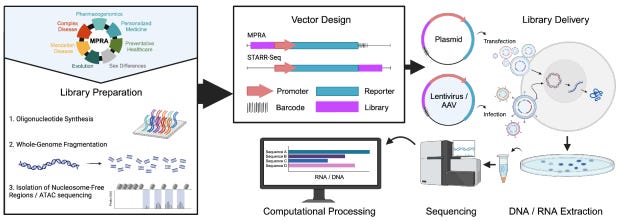
MPRAbase: A Massively Parallel Reporter Assay Database
Science has reached a point where we can measure functional effects for thousands of sequences and variants on gene regulatory activity. There is no comprehensive database of the results of these experiments, but MPRAbase is trying: they have a manually curated set of 129 experiments, with 17 million elements tested across 35 cell types and 4 organisms.
Random Thoughts
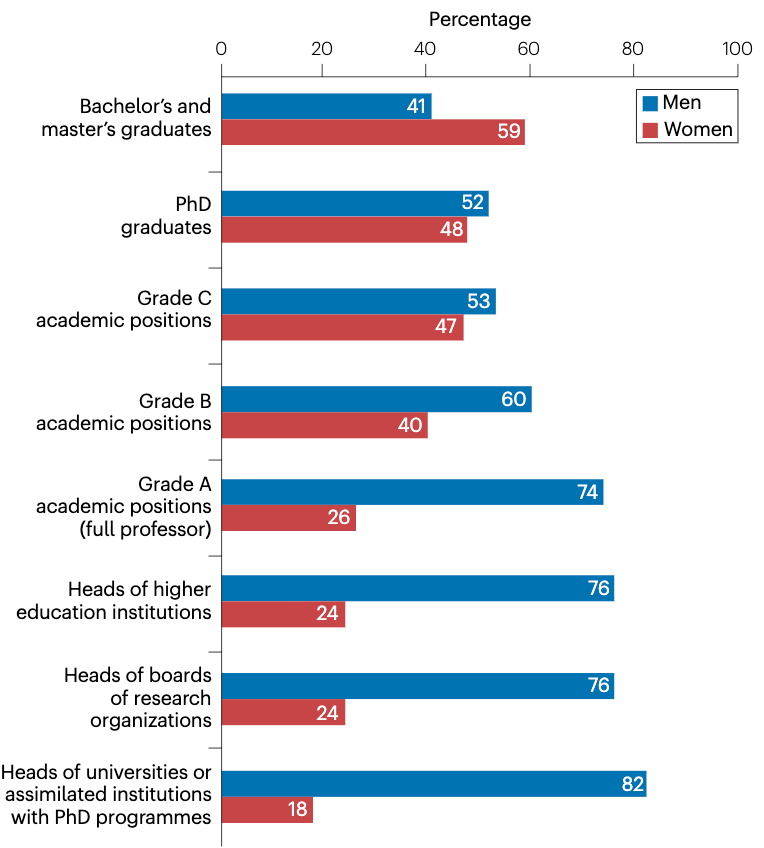
Sexism in Academia is Bad for Science and a Waste of Public Funding
This headline should be no surprise to anyone, but the phenomenon it describes is often under-recognized and seemingly intractable. A huge study of a quarter of a million US academics recently provided weight to the idea that even though women and men graduate at similar rates, academic stages beyond the graduate level are persistently gender-imbalanced because women face extra barriers and end up leaving academia. This article explores the economic consequences of these departures and identifies areas that may have an impact.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
Particle Guidance: non-I.I.D. Diverse Sampling with Diffusion Models by Gabriele Corso
LoGG
A Computational Framework for Solving Wasserstein Lagrangian Flows by Kirill Neklyudov
CARE
Harnessing Geometric Signatures in Causal Representation Learning by Yixin Wang
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋