Portal Weekly #28: a NeurIPS recap of all things TechBio
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on everything happening in TechBio research! This week we’re changing our normal schedule and format. We’ll be doing a recap of NeurIPS-associated papers (that we haven’t previously covered!). Before we jump in, we have a few updates:
🎥 NeurIPS TechBio Social Recap
It was so exciting to see everyone in the community at one of the largest events we’ve ever hosted! We’ll be releasing more photos from the event shortly, in the meantime, here’s a video we got from the venue.
📱 New Account for Portal!
We finally created a Twitter/X account for Portal! Please give us a follow. 🥺
We’ll be using that account to share updates on upcoming LoGG/M2D2/CARE reading groups, the release of recordings, new blogs, new meetups, and more! You can also stay up to date by joining the community directly.
We’ll be off next week for the holidays, but we’ll see you in January!
💬 Upcoming talks
We’ll be off for the holidays, so there are no talks planned for the next week.
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
We chose a cross-section of papers and talks that stood out to us from NeurIPS and associated workshops, but can’t cover it all! Use this as a jumping-off point and check out the links to the workshops at the bottom of the section.
Augmenting Large Language Models With Chemistry Tools
LLMs are popular and have shown good results across many domains, but struggle with chemistry. This study introduces an LLM chemistry agent, augmented with 18 expert-designed tools, that can autonomously plan and execute chemical synthesis and guide the discovery of new chemical matter. Their technical soundness, analysis of limitations and scope of experiments impressed the reviewers.
Holistic Chemical Evaluation Reveals Pitfalls in Reaction Prediction Models
It’s not always intuitive how best to assess out-of-distribution generalization, especially when it comes to chemistry. This study explores generalization in reaction prediction models and introduces ChORISO, a large curated dataset based on previously extracted reactions from high-impact journals. Thinking more carefully about realistic benchmarks suitable for problems of interest will be important for advancement and ensuring that ML makes meaningful contributions.
Prot2Text: Multimodal Protein’s Function Generation with GNNs and Transformers
Most existing methods for predicting a protein’s function treat it as a multi-classification problem. This study proposes prediction in a free text style. They combine graph neural networks and LLMs to integrate diverse data types, assessing the effectiveness of their model on a multimodal protein dataset from SwissProt. Multi-modality is widely expected to be a powerful approach, and stepping away from the classic categorical classifications is also an interesting way to tackle protein function prediction.
DrugImprover: Utilizing Reinforcement Learning for Multi-Objective Alignment in Drug Optimization
Reinforcement learning approaches are increasingly being applied to drug discovery and optimization. This study introduces not only a framework for improving efficiency in drug optimization and an algorithm for finetuning objective-oriented properties, but also a dataset of 1 million compounds with docking scores for cancer-related proteins and SARS-CoV-2 proteins that might be useful for other purposes.
PoseCheck: Generative Models for 3D Structure-Based Drug Design Produce Unrealistic Poses
Multiple studies are finding that the molecules generated by deep learning protein-ligand docking methods often adopt physically unrealistic 3D poses. Both PoseCheck and PoseBusters (we recently had an M2D2 talk on it, check it out) find that state-of-the-art deep learning methods don’t do as well as traditional baselines, and the PoseCheck paper makes recommendations for future research identifying failure modes in structure-based drug design.
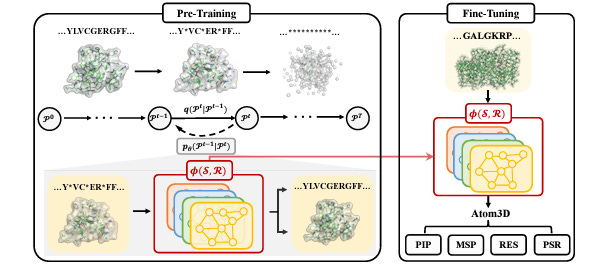
Pre-Training Protein Encoder via Siamese Sequence-Structure Diffusion Trajectory Prediction
Self-supervised pretraining tries to build more universal representations that can then be tailored to the task at hand; most approaches in the protein space either focus on either sequences or structures. This study pretrains a protein encoder to recover sequences and structures along a joint diffusion trajectory. They also attempt to capture correlations between different conformations of a protein. Their models, DiffPreT and SiamDiff, seem to be effective on a range of protein function prediction tasks.
AlphaFold Meets Flow Matching for Generating Protein Ensembles
With recent breakthroughs in protein structure prediction, the next step is predicting structural ensembles. Simultaneously, diffusion models have advanced generative modeling tremendously. This study combines AlphaFold and ESMFold with flow matching, a modern generative modeling framework, to sample the conformational landscape of proteins. The resulting method demonstrates a range of improvements and can predict additional properties depending on how fine-tuned the model is.
Identifying Effects of Disease on Single-Cells with Domain-Invariant Generative Modeling
A core challenge in computational biology is predicting the effects of disease on healthy tissue. This study uses a "single-cell Domain Shift Autoencoder (scDSA)" to separate disease-invariant and disease-specific gene programs at single-cell resolution. Not only does the model learn interaction of disease effects and cell types but it also captures interpretable representations of diseases, which is very useful in a more real-world application like the one discussed in the paper.
HyenaDNA: Long-Range Genomic Sequence Modeling at Single Nucleotide Resolution
Genomic DNA sequences contain a wealth of information, but transformer-based models can only attend to a small portion. This limits their ability to incorporate long-range interactions that are expected to be present in DNA. This study presents a genomic foundation model based on Hyena architecture that more effectively models long-range interactions and scales better than Transformer, using orders of magnitude less parameters and pretraining data. We actually recently hosted a talk on HyenaDNA by the study’s first author - check it out!
ProteinGym: Large-Scale Benchmarks for Protein Fitness Prediction and Design
Predicting the effects of mutations in proteins is critical, for applications ranging from bioengineering to clinical research. Current assessments of ML-based protein fitness prediction are variable and often lacking. This study introduces a large and fairly comprehensive set of benchmarks for protein fitness prediction, including 40 models across zero-shot and supervised settings. Opinions of the paper varied widely among reviewers; put on your science hats and read it for yourself!
ProteinShake: Building Datasets and Benchmarks for Deep Learning on Protein Structures
Improving the rigour and reproducibility of bio-related ML will involve more standardization of datasets, processing, and model evaluation. This paper presents a software package to simplify dataset creation and model evaluation for deep learning on protein structures, and provides a set of pre-processed datasets from the Protein Data Bank (PDB) and AlphaFoldDB. They benchmark prediction tasks associated with each dataset and evaluate model generalization, with an eye towards real-world implications.
There was plenty more interesting research being presented, so check out the following websites for more:
Machine Learning in Structural Biology workshop (scroll down for accepted papers)
Deep Generative Models for Health workshop (the papers are here)
New Frontiers of AI for Drug Discovery and Development workshop (the papers are here)
Causal Representation Learning workshop (the papers are here)
AI4Science workshop (the papers are here)
Generative AI and Biology workshop (the papers are here)
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
LoGG
SE(3)-Stochastic Flow Matching for Protein Backbone Generation by Tara Akhound-Sadegh and Joey Bose
There were no M2D2 or CARE talks this week in the lead-up to the holidays, but you can always catch up on previous recordings on our YouTube channel or the Valence Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋