Portal Weekly #39: reinforcing molecular language models, benchmarking ligand-affinity active learning, and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
📅 TechBio Events
Last week was fun. The Portal team had a chance to work with Nucleate and BitsinBio to organize two TechBio events, one in Toronto and one in Montreal. We had awesome panelists who led discussions on the future of AI-enabled drug discovery and a chance to meet so many of you in the community! There are many more events like this in the future so stay tuned on Portal.
If you want to get involved with Portal to help organize events in your area and support the community blog, please fill out this interest form! We’re looking for volunteers to help grow the community :)
💬 Upcoming talks
M2D2 continues next week with a talk by Austin Tripp, who will describe two projects to improve the evaluation of synthesis planning, a key part of molecule discovery.
Join us live on Zoom on Tuesday, March 19th at 11 am ET. Find more details here.
LoGG continues with a presentation by Omer Bar-Tal who will discuss a framework that uses a pretrained text-to-image diffusion model without any further training or finetuning.
Join us live on Zoom on Monday, March 18th at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
Representing molecules as SMILES strings provides a language-like input to transformer models. LLMs are naturally generative, so it’s a logical step to apply the strategy to molecular design. Two recent papers investigate combining molecular LMs with reinforcement learning to explore the chemical space and try to design better molecules for a purpose:
PromptSMILES: Prompting for Scaffold Decoration and Fragment Linking in Chemical Language Models
This study takes inspiration from the use of prompts to condition language generation, like in GPT models. They take advantage of learned relationships between molecular substructures by providing substructures as a “prompt” to force the final molecule to contain the substructure. Because the model has a tendency to consider the molecule complete after one iteration of this process, they use reinforcement learning (with a similar algorithm to the next study, actually) to fine-tune the model to adapt to iterative prompt-based generation. Their results suggest that this approach works well while being more unified in approach than other strategies.
Evaluation of Reinforcement Learning in Transformer-Based Molecular Design
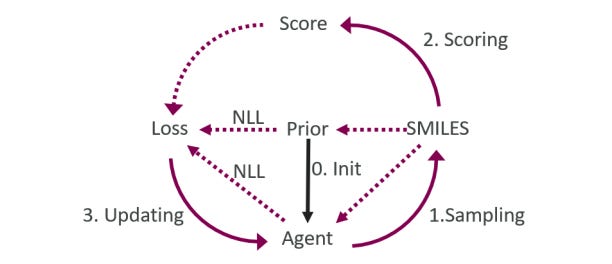
These authors at AstraZeneca previously trained a transformer to generate similar molecules to a given input, adding “property change tokens” to steer the model towards a chemical space of interest. The approach was limited to a preselected set of optimization properties, so they asked whether reinforcement learning could help guide the model more effectively and flexibly. They find, using the REINVENT framework, that they can indeed steer the generation process towards more compounds of interest. They also investigate the impact of pre-trained models, learning steps and rates, giving more insight into different factors affecting performance.
Interested in learning more about large language molecular representation? Amir Barati Farimani gave a talk on the subject at M2D2! If you want to know more about another strategy using chemical language models, Rıza Özçelik gave a talk on chemical language modeling with structured state spaces.
ML for Small Molecules
Benchmarking Active Learning Protocols for Ligand-Binding Affinity Prediction
Active learning (AL) can help us search the vastness of chemical space for desired activities. What effect do different AL parameters and the nature of the dataset have on a model’s performance? This study systematically evaluates the influence of Gaussian Process vs Chemprop models, sample selection protocols, and batch size on overall predictive power and reports trends and takeaways, which may be useful if you’re considering taking advantage of AL advances!
A Small-Molecule TNIK Inhibitor Targets Fibrosis in Preclinical and Clinical Models
It’s nice to see examples of AI-driven approaches speeding up the drug discovery process. This study uses PandaOmics (a commercially-available ML platform that uses transformers and other ML models) to identify targets in fibrosis and generate molecules against the top candidate. They progressed all the way to phase I clinical trials in roughly 18 months. Although we can’t learn too much about the techniques because their ML approach is closed-source, the study does provide insight into the different experimental steps and considerations required to develop a drug candidate.
ML for Atomistic Simulations
Exploring the Frontiers of Condensed-Phase Chemistry with a General Reactive Machine Learning Potential
Machine learning interatomic potentials (MLIPs) have become an efficient alternative to computationally expensive simulations, but often need to be tuned for a specific problem and might not generalize well. This study aims to develop an MLIP applicable to a broad range of reactive chemistry without needing refitting. They automatically sample condensed-phase reactions and use it to study five systems: carbon solid-phase nucleation, graphene ring formation from acetylene, biofuel additives, combustion of methane and the spontaneous formation of glycine from early earth small molecules. They find that their model closely matches experiment or previous studies using traditional methods. It will be interesting to see how general this model is, and if it can be applied to molecular systems!
For more on training ML-based potentials, check out this talk by Stephan Thaler!
ML for Proteins
Protein Language Models are Biased by Unequal Sequence Sampling Across the Tree of Life
Likelihoods from pLMs have been shown to correlate well with protein fitness (catalytic activity, stability, binding affinity etc). This study suggests that there is a species bias in PLMs that affects the likelihoods they output. They found that between 26% and 69% of variance in likelihood could be explained by species identity after controlling for protein type, depending on the pLM. The bias seems to be linked to database makeup, which makes sense (with model organisms overrepresented).
So what does this matter? If we're designing proteins for humans or mice or E.coli (model organisms), it might not matter as much, but they found that unique adaptations of extremophiles (organisms that can tolerate extreme environments like hot springs) might be minimized after pLM likelihood-based design. Something to keep in mind depending on your use-case!
Interested in protein design with large self-supervised models? Kevin K. Yang from Microsoft gave an M2D2 talk covering models that use sequences, structures and biophysical features to generate functional proteins!
ML for Omics
Evaluating the Representational Power of Pre-Trained DNA Language Models for Regulatory Genomics
Studies have suggested that pretrained genomic LMs (gLMs) can get good performance across a broad range of regulatory genomics tasks, but it’s unclear if they have a fundamental understanding of the underlying biology. This study evaluates pretrained gLMs’ ability to predict cell-type-specific functional genomic data on both DNA and RNA levels, without fine-tuning on a specific problem. They argue that this gives a better sense of how well the models have learned fundamental biology. They find that current models’ pretraining does not give clear advantages over more conventional ML approaches. A takeway from the research is that including better inductive biases in gLMs may improve their performance, and that more work is needed until gLMs can be as revolutionary as LLMs have been in other domains.
Open Source
A Compound-Target Pairs Dataset: Differences Between Drugs, Clinical Candidates and Other Bioactive Compounds
There is a limited availability of high-quality open-source data spanning different stages of the drug discovery pipeline. This dataset from EMBL extracts 614,594 compound-target pairs from the open-source bioactivity database ChEMBL, trying to ensure that pairs have either at least one measured activity or are part of the manually curated set of known interactions in ChEMBL (5,109 pairs). Since it is literature-based, the hope is that it can be a higher-confidence resource for open-source drug discovery.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
Removing Biases from Molecular Representations via Information Maximization by Chenyu Wang
Equivariant Scalar Fields for Molecular Docking with Fast Fourier Transforms by Bowen Jing
LoGG
Generalization in diffusion models arises from geometry-adaptive harmonic representation by Zahra Kadkhodaie
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋