Portal Weekly #46: scaling molecular GNNs, retention time prediction for industrial production, CRISPR-GPT, and more.
Join Portal and stay up to date on research happening at the intersection of tech x bio 🧬
Hi everyone 👋
Welcome to another issue of the Portal newsletter where we provide weekly updates on talks, events, and research in the TechBio space!
📅 Upcoming Events
The Machine Learning for Drug Discovery Summer School is now sold out! If you’re looking for other events, check out the 2024 Molecular Machine Learning Conference (MoML) on June 19th. It’s a one-day conference with speakers like Max Jaderberg (Isomorphic Labs), Christine Allen (University of Toronto), Jian Tang (Mila), and more, followed by poster sessions!
If you want to present your work at MoML - the deadline to submit a paper is May 10th at midnight ET. All you need to submit is a 1-pager. Tickets for the conference are also FREE for students!
⭐More details here⭐
💻 Latest Blogs
If a single-cell experiment has some imbalance, does that lead to a loss of biological information when you integrate it with other data? In the latest blog, Hassaan Maan presents the Iniquitate pipeline for assessing imbalanced integration and gives takeaways for representation learning of biological data in general. Check it out!
Interested in sharing your work with the community? Fill out our interest form to get started.
💬 Upcoming talks
M2D2 is off for the summer! It’s been a great season - thank you to all the presenters for the engaging talks and discussions!
LoGG continues next week with a talk by Heli Ben-Hamu, who will talk about a framework for controlling the generation process of diffusion and flow-matching models.
Join us live on Zoom on Monday, May 6th at 11 am ET. Find more details here.
Speakers for CARE talks are usually announced on Fridays after this newsletter goes out. Sign up for Portal and get notifications when a new talk is announced on the CARE events page. You can also follow us on Twitter and LinkedIn!
If you enjoy this newsletter, we’d appreciate it if you could forward it to some of your friends! 🙏
Let’s jump right in!👇
📚 Community Reads
LLMs for Science
CRISPR-GPT: An LLM Agent for Automated Design of Gene-Editing Experiments
Last week we featured a paper in the ML for Proteins section where the authors trained a protein language model on CRISPR genomic machinery, generating new gene editors with increased efficiency and accuracy. This study uses the reasoning abilities of LLMs like GPT-4, Gemini or Claude, enhanced with domain knowledge and external tools, to automate and enhance the design process. They see their method as a bridge between beginner researchers and CRISPR genome engineering techniques.
ML for Small Molecules
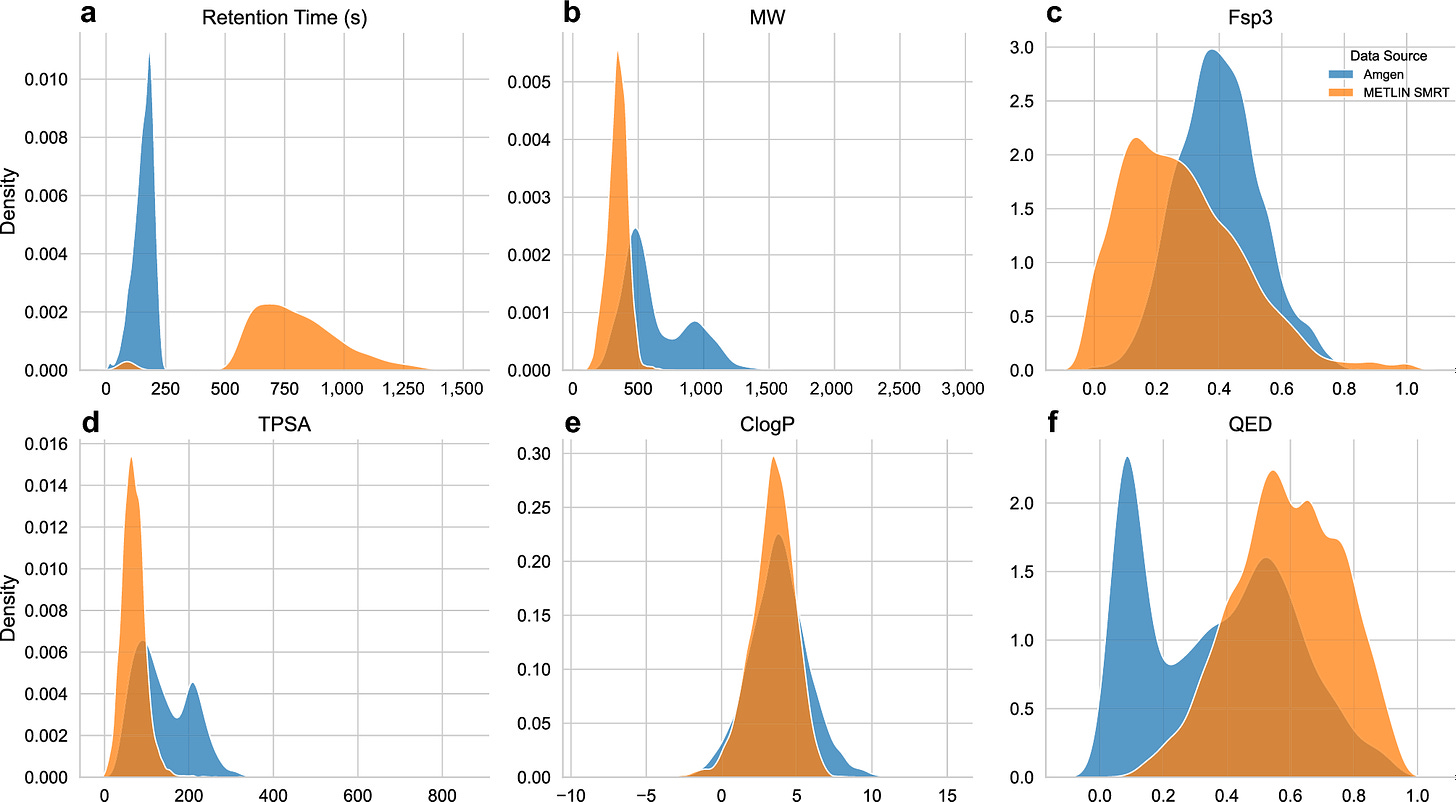
Performance and robustness of small molecule retention time prediction with molecular graph neural networks in industrial drug discovery campaigns
Making sure that you have the right molecule involves using a range of chemical tests and machinery, including chromatography (running molecules through a system that separates the molecules based on how long it takes them to come out; the retention time). This study investigated how machine learning models could predict chromatographic retention time, aiming for robustness in a chemical synthesis production setting. Testing XGBoost, ChemProp and DeepChem models, they found that molecular graph neural networks particularly performed well over time and gave accurate predictions for new chemical series.
Mind Your Prevalence!
Multiple metrics go into assessing and validating the performance of quantitative structure–activity relationship (QSAR) models. This paper presents an analysis of balanced accuracy metrics, particularly MCC (Matthews’ correlation coefficient). It provides a formal, unified framework for understanding prevalence dependence in model validation metrics, and can be useful for those aiming to understand what metrics like MCC actually signify.
Graph Learning
On the Scalability of GNNs for Molecular Graphs
Scaling deep learning models in many domains unlocks emergent behaviour and increased performance. In the molecular domain, GNNs are already intuitive and powerful ways of representing and acting on molecules, but because of the different structural design choices of GNNs, it has been unclear what benefits scaling them actually gives. This study analyzes the effect of scaling depth, width, number of molecules, number of labels, and the diversity in the pretraining datasets, and presents a new graph foundation model, MolGPS, that outperforms the previous state-of-the-art on 26/38 downstream tasks.
Interested in how deep learning models scale in chemistry? Nathan Frey recently wrote a blog post on neural scaling of deep chemical models!
ML for Atomistic Simulations
AutoDock-SS: AutoDock for Multiconformational Ligand-Based Virtual Screening
Ligand-based virtual screening (LBVS) can be useful for identifying new candidate drugs, but current methods may not be able to consider ligand conformational flexibility. This study presents a way of optimizing conformations dynamically based on the reference ligand, which likely gives a more accurate representation. The authors show that their method does better than alternative LBVS methods on the well-known DUD-E and DUD-E+ datasets.
GōMartini 3: From large conformational changes in proteins to environmental bias corrections
Coarse-grained modeling approaches can allow for a less granular but longer time-scale view of molecular dynamics. This paper introduces the latest version of the GōMartini model, which tries to balance computational efficiency and the ability to study proteins in different biological environments. The authors say that this implementation has been extensively tested by the community since the release of the new version of Martini, and that their model can address recent inaccuracies reported in the Martini protein model.
ML for Proteins
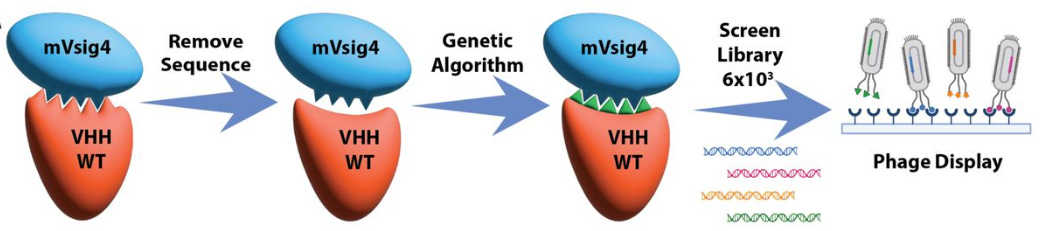
Using antibodies for medicine is limited by the difficulty of designing novel antibodies to bind a specific epitope on a target (the part that is recognized by the immune system). Two recent studies report making headway on the antibody design problem. Atomically accurate de novo design of single-domain antibodies, from the Baker lab, shows that a fine-tuned RFDiffusion can design antibody variable heavy chains (VHHs) that bind user-specified epitopes, and they experimentally confirm binders to four disease-relevant epitopes in the lab. De novo design of high-affinity single-domain antibodies uses an empirical force field FoldX to optimize VHH stability and affinity for original targets, or design VHHs for new targets, reaching low nanomolar affinity in a single design cycle.
Atomically accurate de novo design of single-domain antibodies
De novo design of high-affinity single-domain antibodies
ML for Omics
Inferring gene regulatory networks from single-cell multiome data using atlas-scale external data
Gene regulatory networks (GRNs) are collections of molecular regulators that interact with each other and determine gene activation and silencing in specific cellular contexts. We have better ways of measuring what DNA is read and how much mRNA is around, but learning complex mechanisms of regulation is still difficult. This study presents an approach for inferring GRNs from single-cell paired gene expression and chromatin accessibility data. The authors suggest that this method gives 4- to 7-fold better accuracy in predicting regulatory networks and even allows estimating transcription factor activity just from bulk gene expression data.
Open Source
md-agent - molecular simulation with an agent
MD-Agent is a LLM-agent based toolset for Molecular Dynamics. It's built using Langchain and uses a collection of tools to set up and execute molecular dynamics simulations, particularly in OpenMM. An OpenAI key is necessary for this project.
Congrats to Daniel Wigh for getting ORDerly: Data Sets and Benchmarks for Chemical Reaction Data published in the Journal of Chemical Information and Modeling! We covered his paper when it was on OpenReview back in October, after he brought it to our attention by posting about his work on Portal.
Think we missed something? Join our community to discuss these topics further!
🎬 Latest Recordings
M2D2
EquiReact: An Equivariant Neural Network for Chemical Reactions by Puck van Gerwen
LoGG
Multimodal language models for mapping the genotype-phenotype relationship by Farhan Khodaee
You can always catch up on previous recordings on our YouTube channel or the Portal Events page!
Portal is the home of the TechBio community. Join here and stay up to date on the latest research, expand your network, and ask questions. In Portal, you can access everything that the community has to offer in a single location:
M2D2 - a weekly reading group to discuss the latest research in AI for drug discovery
LoGG - a weekly reading group to discuss the latest research in graph learning
CARE - a weekly reading group to discuss the latest research in causality
Blogs - tutorials and blog posts written by and for the community
Discussions - jump into a topic of interest and share new ideas and perspectives
See you at the next issue! 👋